Overview
epifitter offers three complementary fitting
workflows:
-
fit_lin() for linearized model comparison;
-
fit_nlin() for nonlinear fitting of two-parameter
models;
-
fit_nlin2() for nonlinear fitting when the asymptote
K should be estimated.
Compare models quickly with fit_lin()
set.seed(1)
epi <- sim_logistic(N = 60, y0 = 0.01, dt = 5, r = 0.12, alpha = 0.2, n = 4)
fit_lin_out <- fit_lin(time = epi$time, y = epi$random_y)
knitr::kable(fit_lin_out$stats_all, digits = 4)
| 1 |
Logistic |
0.1202 |
0.0011 |
0.1180 |
0.1224 |
-4.5730 |
0.0394 |
0.9957 |
0.1504 |
0.9979 |
0.0102 |
0.0095 |
0.0111 |
| 2 |
Gompertz |
0.0694 |
0.0022 |
0.0649 |
0.0739 |
-1.9893 |
0.0792 |
0.9505 |
0.3021 |
0.9746 |
0.0007 |
0.0002 |
0.0020 |
| 3 |
Exponential |
0.0780 |
0.0029 |
0.0722 |
0.0839 |
-4.0617 |
0.1032 |
0.9346 |
0.3938 |
0.9662 |
0.0172 |
0.0140 |
0.0212 |
| 4 |
Monomolecular |
0.0422 |
0.0028 |
0.0366 |
0.0478 |
-0.5113 |
0.0983 |
0.8215 |
0.3751 |
0.9020 |
-0.6674 |
-1.0315 |
-0.3687 |
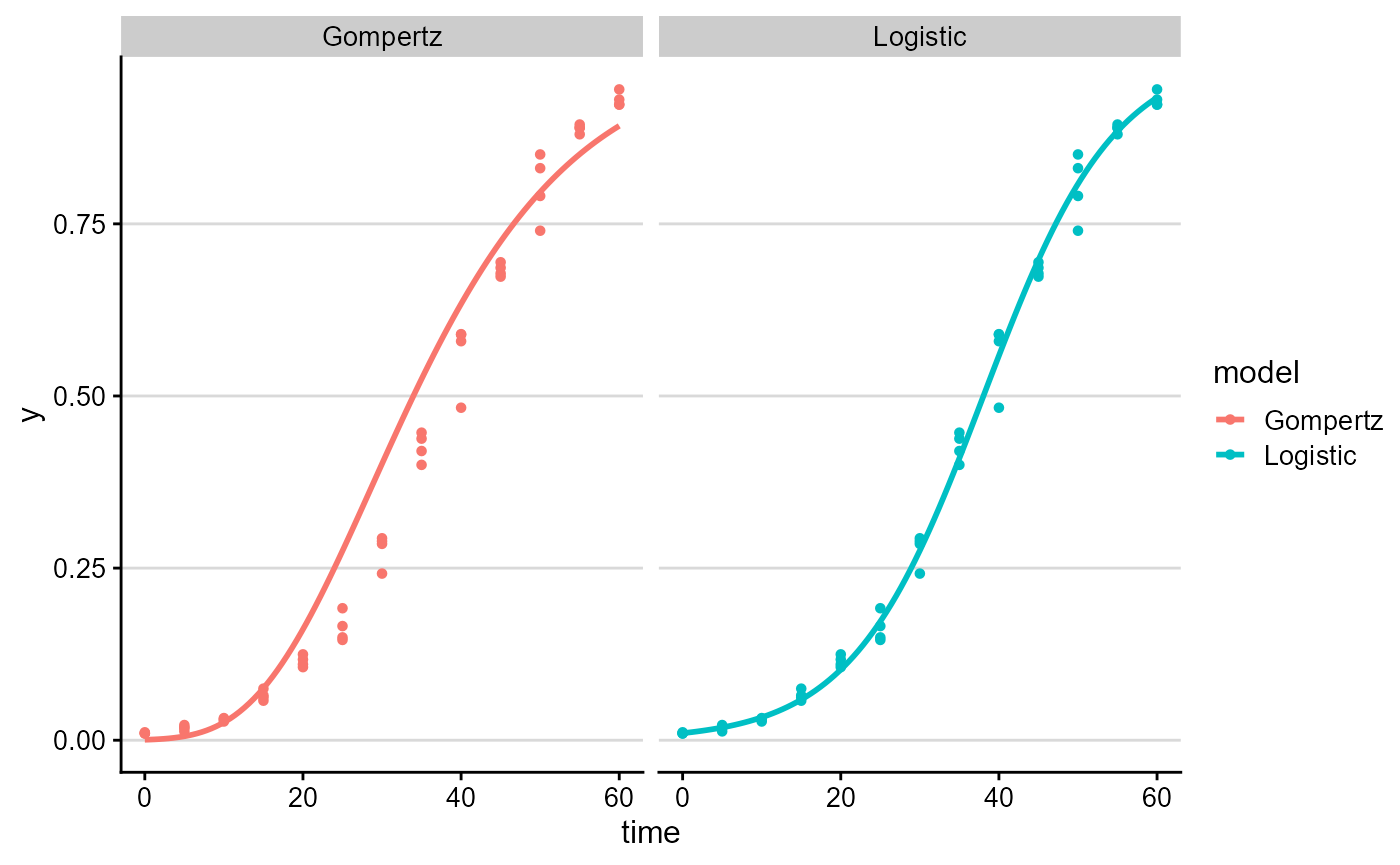
plot_fit(fit_lin_out, models = c("Logistic", "Gompertz"))
Nonlinear fitting with starting values
fit_nlin_out <- fit_nlin(
time = epi$time,
y = epi$random_y,
starting_par = list(y0 = 0.01, r = 0.03)
)
fit_nlin_out$stats_all
Estimate K when the epidemic plateaus below 1
epi_partial <- epi %>%
mutate(random_y = random_y * 0.8)
fit_k <- fit_nlin2(
time = epi_partial$time,
y = epi_partial$random_y,
starting_par = list(y0 = 0.01, r = 0.03, K = 0.7)
)
fit_k$stats_all
Grouped fitting with fit_multi()
epi1 <- sim_gompertz(N = 50, y0 = 0.001, dt = 5, r = 0.08, alpha = 0.2, n = 3)
epi2 <- sim_gompertz(N = 50, y0 = 0.002, dt = 5, r = 0.11, alpha = 0.2, n = 3)
multi_epi <- bind_rows(epi1, epi2, .id = "curve")
multi_fit <- fit_multi(
time_col = "time",
intensity_col = "random_y",
data = multi_epi,
strata_cols = "curve"
)
knitr::kable(head(multi_fit$Parameters), digits = 4)
| 1 |
1 |
Gompertz |
0.0813 |
0.0016 |
0.0780 |
0.0846 |
-1.9382 |
0.0480 |
0.9878 |
0.1475 |
0.9939 |
0.0010 |
0.0005 |
0.0018 |
| 1 |
2 |
Logistic |
0.1623 |
0.0091 |
0.1438 |
0.1808 |
-5.1753 |
0.2685 |
0.9116 |
0.8244 |
0.9537 |
0.0056 |
0.0033 |
0.0097 |
| 1 |
3 |
Monomolecular |
0.0449 |
0.0026 |
0.0396 |
0.0502 |
-0.3618 |
0.0769 |
0.9058 |
0.2360 |
0.9506 |
-0.4359 |
-0.6796 |
-0.2276 |
| 1 |
4 |
Exponential |
0.1174 |
0.0111 |
0.0948 |
0.1400 |
-4.8135 |
0.3276 |
0.7838 |
1.0058 |
0.8788 |
0.0081 |
0.0042 |
0.0158 |
| 2 |
1 |
Gompertz |
0.1126 |
0.0015 |
0.1095 |
0.1157 |
-1.8561 |
0.0448 |
0.9944 |
0.1377 |
0.9972 |
0.0017 |
0.0009 |
0.0029 |
| 2 |
2 |
Monomolecular |
0.0802 |
0.0035 |
0.0729 |
0.0874 |
-0.5657 |
0.1050 |
0.9427 |
0.3224 |
0.9705 |
-0.7606 |
-1.1810 |
-0.4212 |
knitr::kable(head(multi_fit$Data), digits = 4)
| 1 |
0 |
0.0011 |
Exponential |
-6.8418 |
0.0081 |
-0.0071 |
| 1 |
0 |
0.0011 |
Monomolecular |
0.0011 |
-0.4359 |
0.4370 |
| 1 |
0 |
0.0011 |
Logistic |
-6.8408 |
0.0056 |
-0.0046 |
| 1 |
0 |
0.0011 |
Gompertz |
-1.9231 |
0.0010 |
0.0001 |
| 1 |
5 |
0.0076 |
Exponential |
-4.8831 |
0.0146 |
-0.0070 |
| 1 |
5 |
0.0076 |
Monomolecular |
0.0076 |
-0.1473 |
0.1549 |

