git clone https://github.com/AlvesKS/paper_ENSO_SBR_damage.git
cd your-repoIntroduction
This repository contains the code and data processing workflow supporting the analyses presented in the study:
“Quantifying ENSO-mediated shifts in soybean rust impact: Yield loss dynamics and management implications in Brazil.”
Soybean rust (SBR), caused by Phakopsora pachyrhizi, is a major threat to soybean production in Brazil. The study investigates how climate variability, specifically the El Niño Southern Oscillation (ENSO), influences SBR-related yield losses and the performance of disease management strategies. To this end, we analyzed a comprehensive dataset of 417 field trials conducted across 73 locations in Brazil from 2005 to 2020.
The primary goals of the analysis were to:
- Estimate disease damage coefficients (i.e., the relationship between disease severity and yield loss) under different ENSO phases (warm, Neutral, cold).
- Assess the effectiveness of control measures across these climatic contexts.
- Quantify how ENSO conditions modulate both disease impact and management outcomes.
This codebase includes all steps for:
- Cleaning and structuring raw trial data
- Classifying ENSO phases
- Fitting statistical models to estimate damage coefficients
- Performing meta-analyses and producing summary figures and tables
Getting started
This project uses renv to manage package dependencies and ensure reproducible computational environments. Follow the steps below to set up and execute the code.
Prerequisites
R (≥ 4.4.1) Ensure you have a compatible version of R installed. You can download R from CRAN.
Git (optional but recommended) To clone the repository, install Git from git-scm.com. Alternatively, you can download the repository as a ZIP.
Project Files
Steps to Initialize the Environment
- Clone or Download the Repository
Alternatively, you can download the repository as a ZIP file and extract it to your working directory using the code below:
download.file("https://github.com/AlvesKS/paper_ENSO_SBR_damage/archive/refs/heads/main.zip", destfile = "paper-white-mold-prediction-modeling.zip", mode = "wb")
unzip("paper-white-mold-prediction-modeling.zip", list = FALSE)- Open the Project in R
The repository files should be inside the paper-white-mold-prediction-modeling/ directory, which should be locatated in your working directory. Open the file .Rproj file.
- Install or update the
renvpackage (if not already installed)
If you haven’t installed renv yet, run the following command in your R console. If you already have renv, ensure it is updated to the latest version.
install.packages("renv")- Restore the Project Library
- This command reads the renv.lock file and installs the exact package versions into a project-specific library (
renv/library/).
- If prompted to update renv itself, follow the message and restart R afterward.
renv::restore()- Verify Successful Restoration
After renv::restore() completes, you should see messages indicating that required packages were installed. You can check the status with:
renv::status()If all dependencies are up to date, renv::status() will report no divergences from the lockfile.
Running the Analysis
- Locate the Main Script
The primary analysis script are located in the R/ directory named as main_damage_coef.qmd.
- Execute the Workflow
From now on, you can run the analysis by executing the code chunks in the R quarto file. The code is organized into sections, and you should each execute section sequentialy or knit the entire document to generate a report.
- Inspect and Export Outputs
Output figures (PNG/PDF) are saved in the figs/ folder.
Packages
library(tidyverse)
library(gsheet)
library(cowplot)
library(patchwork)
library(lemon)
library(lme4)
library(ggforce)
library(ggrepel)
library(lmerTest)
library(emmeans)
library(multcomp)
library(ggthemes)
library(metafor)
library(minpack.lm)theme_set(theme_half_open(font_size = 10))Data
Soybean Rust data
data_load = read.csv("data/raw-data-2005-2020.csv") |>
mutate(sev = as.numeric(sev),
yld = as.numeric(yld))Warning: There were 2 warnings in `mutate()`.
The first warning was:
ℹ In argument: `sev = as.numeric(sev)`.
Caused by warning:
! NAs introduzidos por coerção
ℹ Run `dplyr::last_dplyr_warnings()` to see the 1 remaining warning.ENSO data
enso_data = read.csv("data/enso.csv")Classifying the years based on Oceanic Niño Index (ONI) on the October, November, and December (OND) trimester. Seasons with ONI higher then 75 percentiles were classified as warm, year with ONI lower then its 25 percentiles were classified as Cold, and the years with ONI within the 25 and 75 percentiles were classified as neutral.
enso_data_class = enso_data |>
mutate(year = as.character(Year+1)) |>
dplyr::select(-Year) |>
mutate(selected_trimester = OND) |>
dplyr::select(selected_trimester, year) |>
filter(!is.na(selected_trimester),
year != 2021) |>
mutate(quantile_0.75 = quantile(selected_trimester, 0.75),
quantile_0.25 = quantile(selected_trimester, 0.25)) |>
mutate(enso = case_when(selected_trimester > quantile_0.75 ~ "Warm",
selected_trimester <=quantile_0.25 ~ "Cold",
selected_trimester <= quantile_0.75 & selected_trimester > quantile_0.25 ~ "Neutral"),
year = as.numeric(year))
enso_data_classenso_data_class |>
summarise(quantile_0.75 = unique(quantile_0.75),
quantile_0.25 = unique(quantile_0.25))enso_gg = enso_data_class|>
mutate(enso = factor(enso, levels = c("Neutral", "Warm", "Cold")))|>
ggplot(aes( as.factor(year),selected_trimester, color = enso))+
geom_hline(yintercept = 0)+
geom_hline(yintercept = c(-0.725,0.75), linetype = 2, color = "gray")+
geom_errorbar(aes(ymin=0, ymax = selected_trimester), width = 0, color = "black")+
geom_point(size = 3)+
scale_color_colorblind()+
theme_half_open(font_size = 12)+
scale_y_continuous(breaks = seq(-3,3,by = 0.75), limits = c(-3,3))+
theme(axis.text.x = element_text(angle = 45, hjust = 1),
# legend.position = "top",
strip.background = element_blank())+
labs(x = "Crop Season",
y = "Oceanic Niño Index",
color = "ENSO phase (OND)")
enso_gg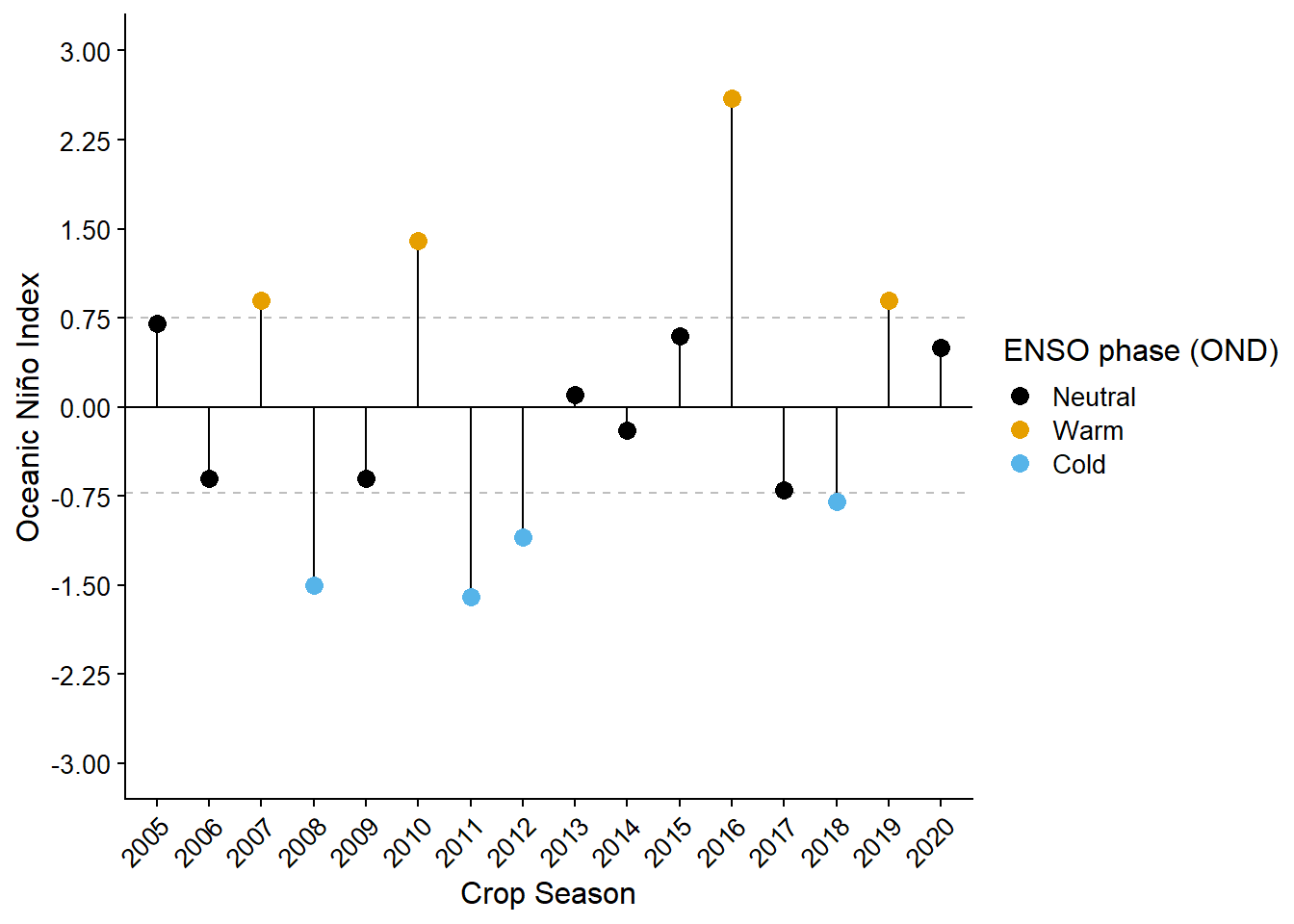
ggsave("figs/ONI_OND.png", dpi = 600, height = 4, width = 7, bg = "white")Counting the number of year for each phase
enso_data_class |>
count(enso)Data wrangling
data_load2 = data_load |>
mutate(sev = as.numeric(sev)) |>
full_join(enso_data_class) |>
mutate(enso = factor(enso, levels = c("Neutral", "Warm", "Cold"))) |>
filter(!is.na(sev),
!is.na(yld)) |>
mutate(study = as.factor(study)) |>
mutate(region = case_when(state %in% c("SP","BA","MG", "MS", "MT", "GO", "MA", "DF", "TO")~"North",
state %in% c("RS","SC","PR") ~"South"),
region =factor(region, levels = c("South","North"))) |>
group_by(study) |>
mutate(difer = max(sev) - min(sev)) |>
filter(difer>5)Joining with `by = join_by(year)`data_load2 |>
filter(active_ingred == "CHECK") |>
ggplot(aes(sev, difer))+
geom_point(color = "black", size =2, shape =1)+
geom_hline(yintercept = 5, linetype = "dashed", color = "gray", size = 1.2)+
geom_abline(slope = 1, intercept = 0, color = "darkred", size = 1.4)+
annotate("text",x = 100, y = 5, label = "Minimum difference = 5 p.p", vjust = -1, hjust = 1)+
coord_cartesian(xlim = c(0,100),
ylim = c(0,100))Warning: Using `size` aesthetic for lines was deprecated in ggplot2 3.4.0.
ℹ Please use `linewidth` instead.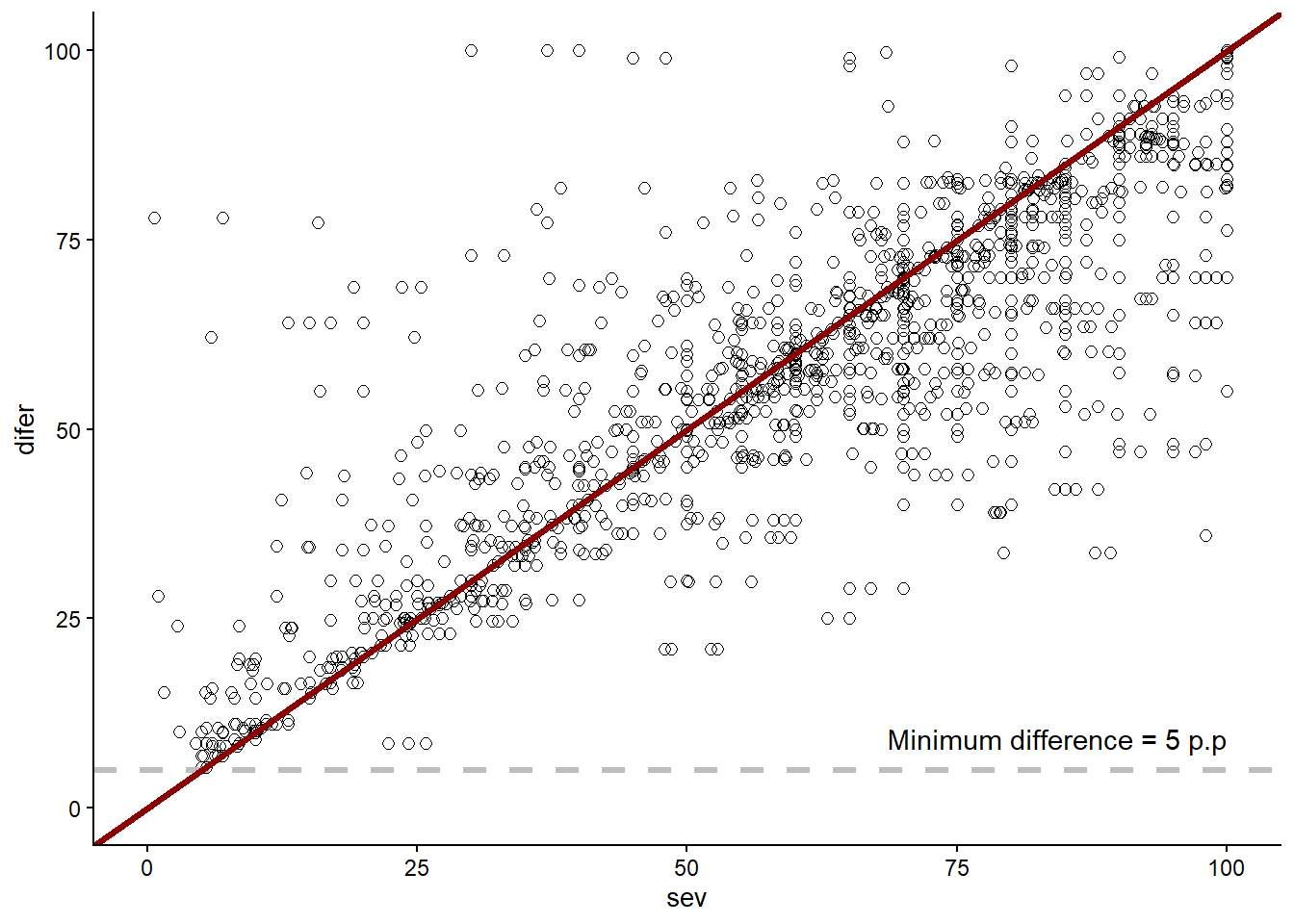
unique(data_load2$state) [1] "BA" "PR" "MG" "MS" "MT" "GO" "SP" "MA" "DF" "TO" "RS" "SC"length(unique(data_load2$state))[1] 12data_load2 |>
group_by(state) |>
summarise(n_loc = length(unique(location))) |>
summarise(sum(n_loc))length(unique(data_load2$year))[1] 16data_load2 |>
group_by(enso) |>
summarise(n_study = length(unique(study)),
n_year = length(unique(year)),
n_loc = length(unique(location)),
n_state = length(unique(state)))Tranformations
Converting percent severity into proportion and calculating logits
only_check_df = data_load2 |>
filter(active_ingred == "CHECK") |>
mutate(sev = case_when(sev == 100 ~ 99.9,
sev == 0 ~ 0.1,
sev>0 & sev<100 ~ sev),
logit_sev = DescTools::Logit(sev/100))Exploratory analysis
Average, min and max severity
Overal
data_load2 |>
filter(active_ingred == "CHECK") |>
ungroup() |>
summarise(avg_sev = mean(sev),
max_sev = max(sev),
min_sev = min(sev))by year
data_load2 |>
filter(active_ingred == "CHECK") |>
group_by(year) |>
summarise(avg_sev = mean(sev),
max_sev = max(sev),
min_sev = min(sev)) |>
arrange(max_sev)By ENSO state
data_load2 |>
filter(active_ingred == "CHECK") |>
group_by(enso) |>
summarise(avg_sev = mean(sev),
max_sev = max(sev),
min_sev = min(sev))plot
enso_sev_gg = data_load2 |>
filter(active_ingred == "CHECK") |>
ggplot(aes(enso, sev, color = enso))+
geom_sina(color = "gray")+
geom_boxplot(fill= NA, size = 1)+
theme_half_open(font_size = 12)+
scale_color_colorblind()+
labs( y = "",
x = "",
color = "",
title = "")
enso_sev_gg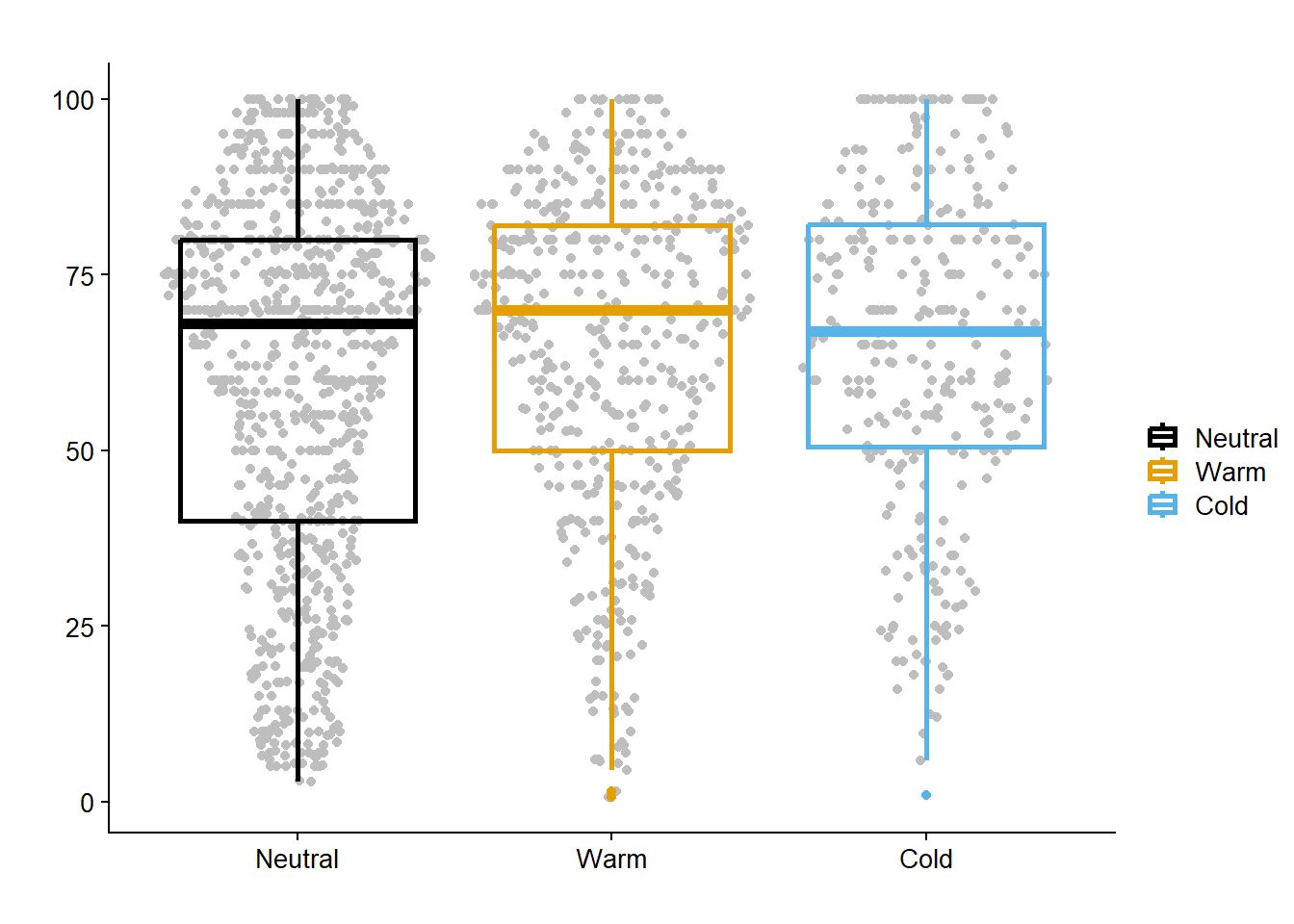
Graph over time
sev_untreat_gg = only_check_df |>
ggplot(aes(as.factor(year), sev, color = enso))+
geom_sina(color = "gray80")+
geom_boxplot(fill =NA, size = 1, outlier.color = NA)+
labs( y = "Severity (%)",
x = "",
color = "",
title = " ")+
theme_half_open(font_size = 12)+
# facet_wrap(~region)+
scale_color_colorblind()
sev_untreat_gg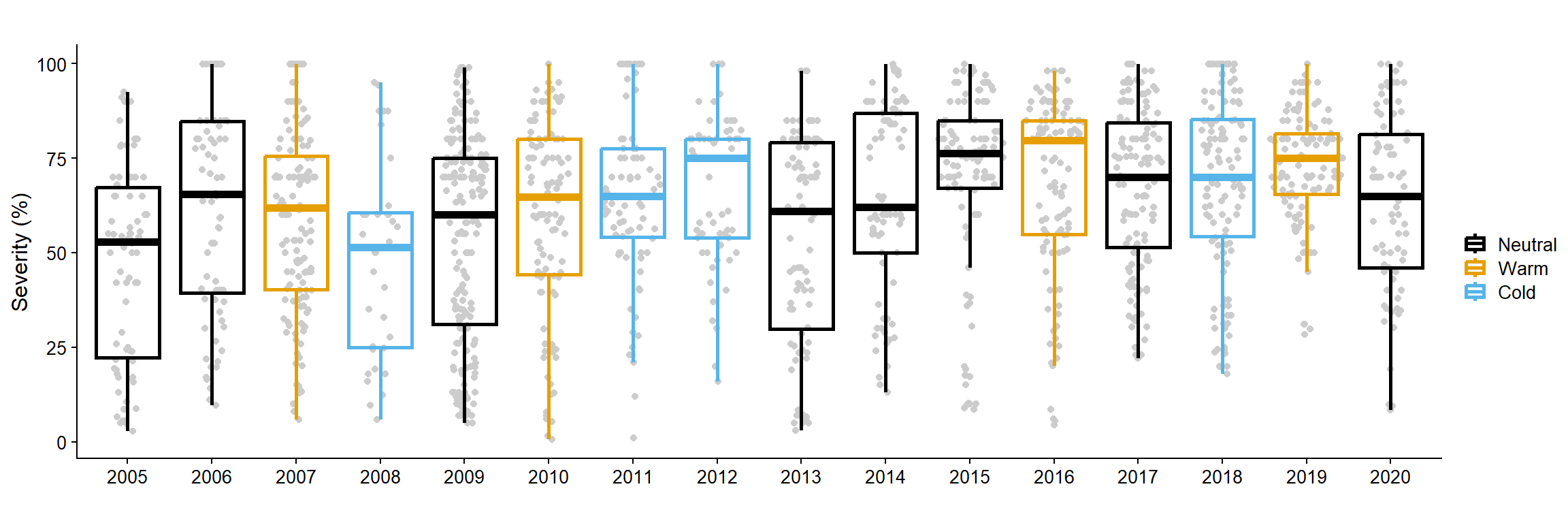
Average, min and max yield
Overal
data_load2 |>
filter(active_ingred == "CHECK") |>
ungroup() |>
summarise(avg_yld = mean(yld),
max_yld = max(yld),
min_yld = min(yld))by year
data_load2 |>
filter(active_ingred == "CHECK") |>
group_by(year) |>
summarise(avg_yld = mean(yld),
max_yld = max(yld),
min_yld = min(yld)) |>
arrange(avg_yld)By ENSO state
data_load2 |>
filter(active_ingred == "CHECK") |>
group_by(enso)|>
summarise(avg_yld = mean(yld),
max_yld = max(yld),
min_yld = min(yld)) |>
arrange(avg_yld)Modeling Severity (untreated) and ENSO
Mixed-effect model
model_check = lmer(logit_sev ~ enso+ (1|year/study), data = only_check_df,REML = F)Model summary
summary(model_check)Linear mixed model fit by maximum likelihood . t-tests use Satterthwaite's
method [lmerModLmerTest]
Formula: logit_sev ~ enso + (1 | year/study)
Data: only_check_df
AIC BIC logLik deviance df.resid
4506.7 4539.3 -2247.3 4494.7 1680
Scaled residuals:
Min 1Q Median 3Q Max
-7.5458 -0.2094 0.0000 0.2012 9.5043
Random effects:
Groups Name Variance Std.Dev.
study:year (Intercept) 2.80243 1.6740
year (Intercept) 0.07201 0.2683
Residual 0.35208 0.5934
Number of obs: 1686, groups: study:year, 418; year, 16
Fixed effects:
Estimate Std. Error df t value Pr(>|t|)
(Intercept) 0.6678 0.1523 13.0217 4.385 0.000735 ***
ensoWarm 0.2028 0.2544 11.8761 0.797 0.441117
ensoCold 0.4745 0.2843 16.4863 1.669 0.113967
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation of Fixed Effects:
(Intr) ensWrm
ensoWarm -0.599
ensoCold -0.536 0.321Pairwise comparison
pairs(emmeans(model_check, ~enso)) contrast estimate SE df t.ratio p.value
Neutral - Warm -0.203 0.290 18.6 -0.699 0.7671
Neutral - Cold -0.474 0.320 23.0 -1.485 0.3165
Warm - Cold -0.272 0.355 21.4 -0.764 0.7283
Degrees-of-freedom method: kenward-roger
P value adjustment: tukey method for comparing a family of 3 estimates All Severity data (plot)
sev_gg = data_load2 |>
ggplot(aes(as.factor(year), sev, color = enso))+
geom_sina(color = "gray80")+
geom_boxplot(fill =NA, size = 1, outlier.color = NA)+
labs( y = "Severity (%)",
x = "Growing season",
color = "",
title = "Data from all plots")+
theme_half_open(font_size = 12)+
# facet_wrap(~region)+
scale_color_colorblind()
sev_gg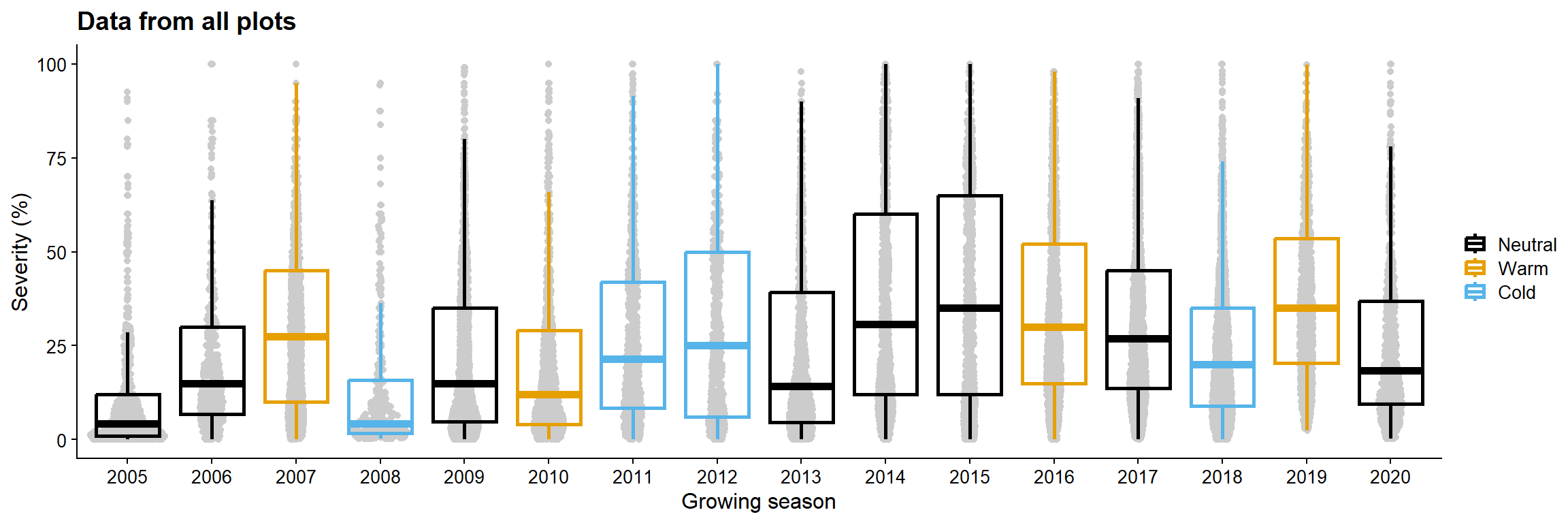
Modeling soybean yield and ENSO
enso_yld_gg = data_load2 |>
filter(active_ingred == "CHECK") |>
ggplot(aes(enso, yld, color = enso))+
geom_sina(color = "gray")+
geom_boxplot(fill= NA, size = 1)+
scale_color_colorblind()+
theme_half_open(font_size = 12)+
labs( y = "",
x = "",
color = "",
title = "")
enso_yld_gg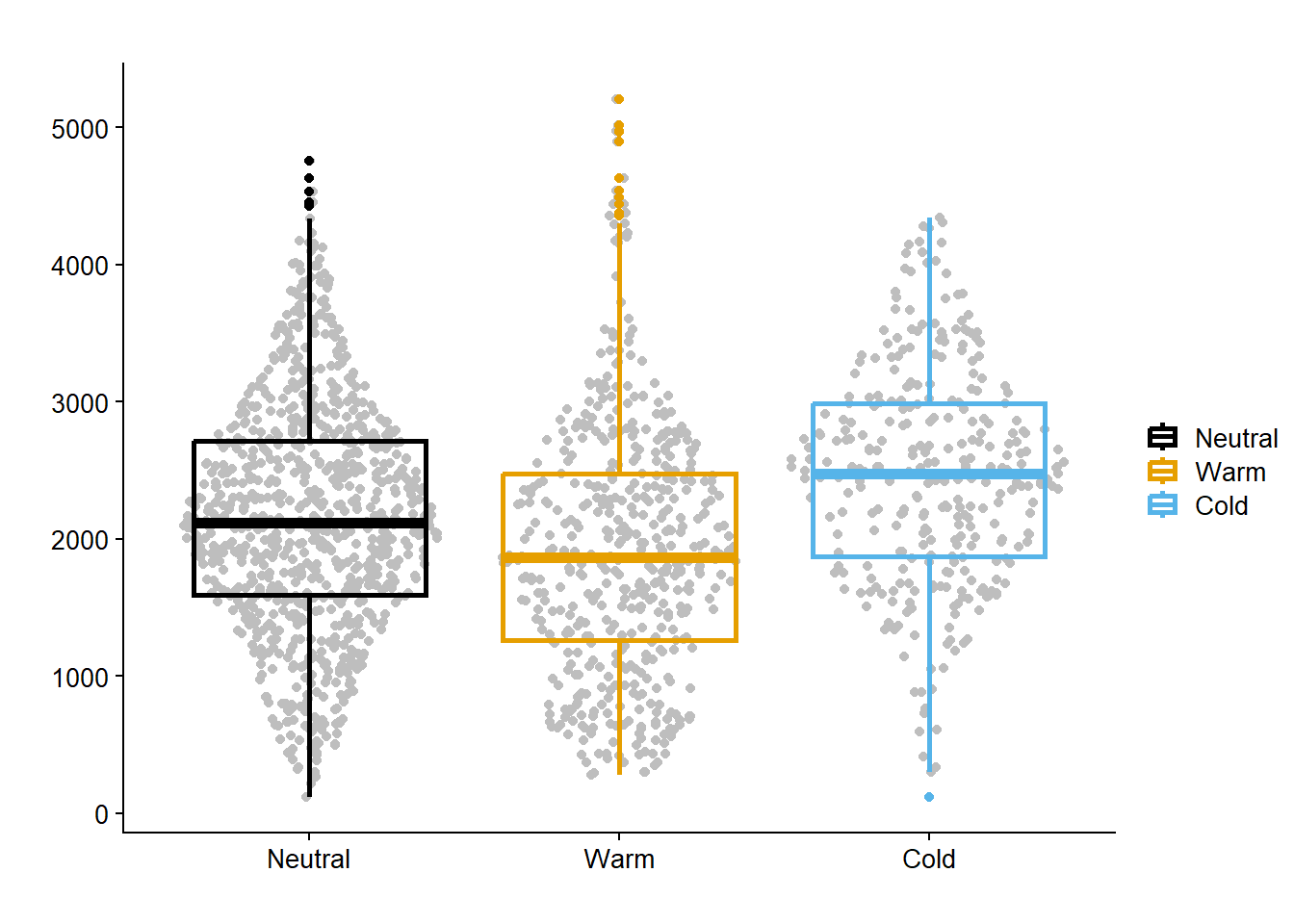
Graph over time
yld_untreat_gg = data_load2 |>
filter(active_ingred == "CHECK")|>
ggplot(aes(as.factor(year), yld, color = enso))+
geom_sina(color = "gray80")+
geom_boxplot(size =1, fill = NA, outlier.colour = NA)+
labs( y = "Yield (kg/ha)",
x = "Growing season",
color = "",
title = "")+
theme_half_open(font_size = 12)+
scale_color_colorblind()
yld_untreat_gg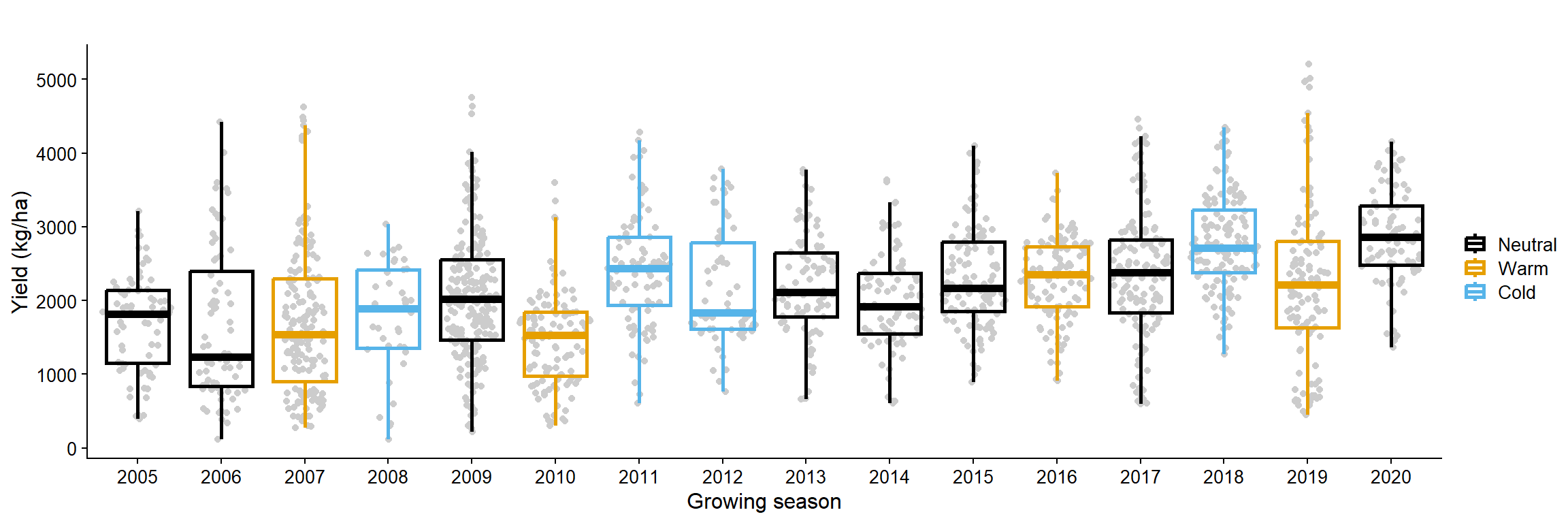
Mixed-effect model
model_check_yld = lmer(yld ~ enso + (1|year/study), data = only_check_df, REML = F)Model summary
summary(model_check_yld)Linear mixed model fit by maximum likelihood . t-tests use Satterthwaite's
method [lmerModLmerTest]
Formula: yld ~ enso + (1 | year/study)
Data: only_check_df
AIC BIC logLik deviance df.resid
24878.7 24911.3 -12433.4 24866.7 1680
Scaled residuals:
Min 1Q Median 3Q Max
-4.5778 -0.4421 0.0067 0.4548 5.2378
Random effects:
Groups Name Variance Std.Dev.
study:year (Intercept) 618988 786.8
year (Intercept) 96443 310.6
Residual 57242 239.3
Number of obs: 1686, groups: study:year, 418; year, 16
Fixed effects:
Estimate Std. Error df t value Pr(>|t|)
(Intercept) 2148.18 123.74 14.70 17.361 3.36e-11 ***
ensoWarm -219.25 211.23 13.94 -1.038 0.317
ensoCold 181.70 221.18 16.40 0.822 0.423
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation of Fixed Effects:
(Intr) ensWrm
ensoWarm -0.586
ensoCold -0.559 0.328Pairwise comparison
cld(emmeans(model_check_yld, ~enso))All yield data
Graph over time
yld_gg = data_load2 |>
ggplot(aes(as.factor(year), yld, color = enso))+
geom_sina(color = "gray80")+
geom_boxplot(size =1, fill = NA, outlier.colour = NA)+
labs( y = "Yield (kg/ha)",
x = "Growing season",
color = "",
title = "Data from all plots")+
theme_half_open(font_size = 12)+
scale_color_colorblind()
yld_gg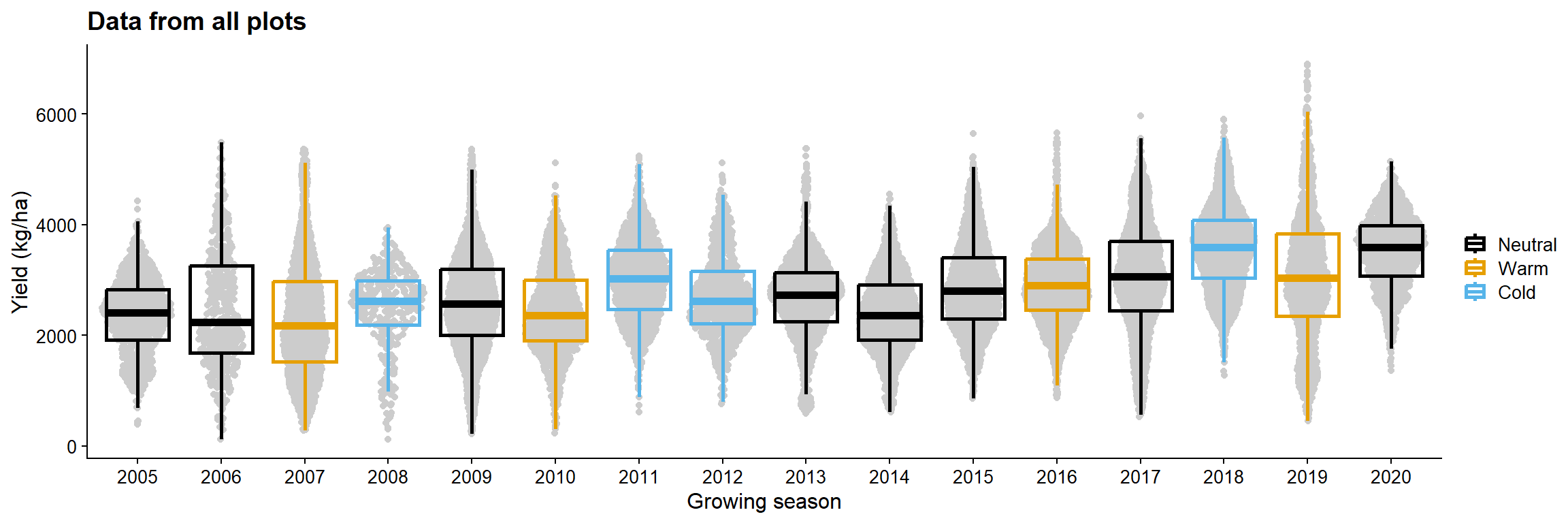
Combo plot
sev_untreat_gg + enso_sev_gg+
yld_untreat_gg +enso_yld_gg+
plot_annotation(tag_levels = "a")+
plot_layout(ncol = 2,
widths = c(1,0.25),
guides = "collect")&
theme(axis.text = element_text(size =8))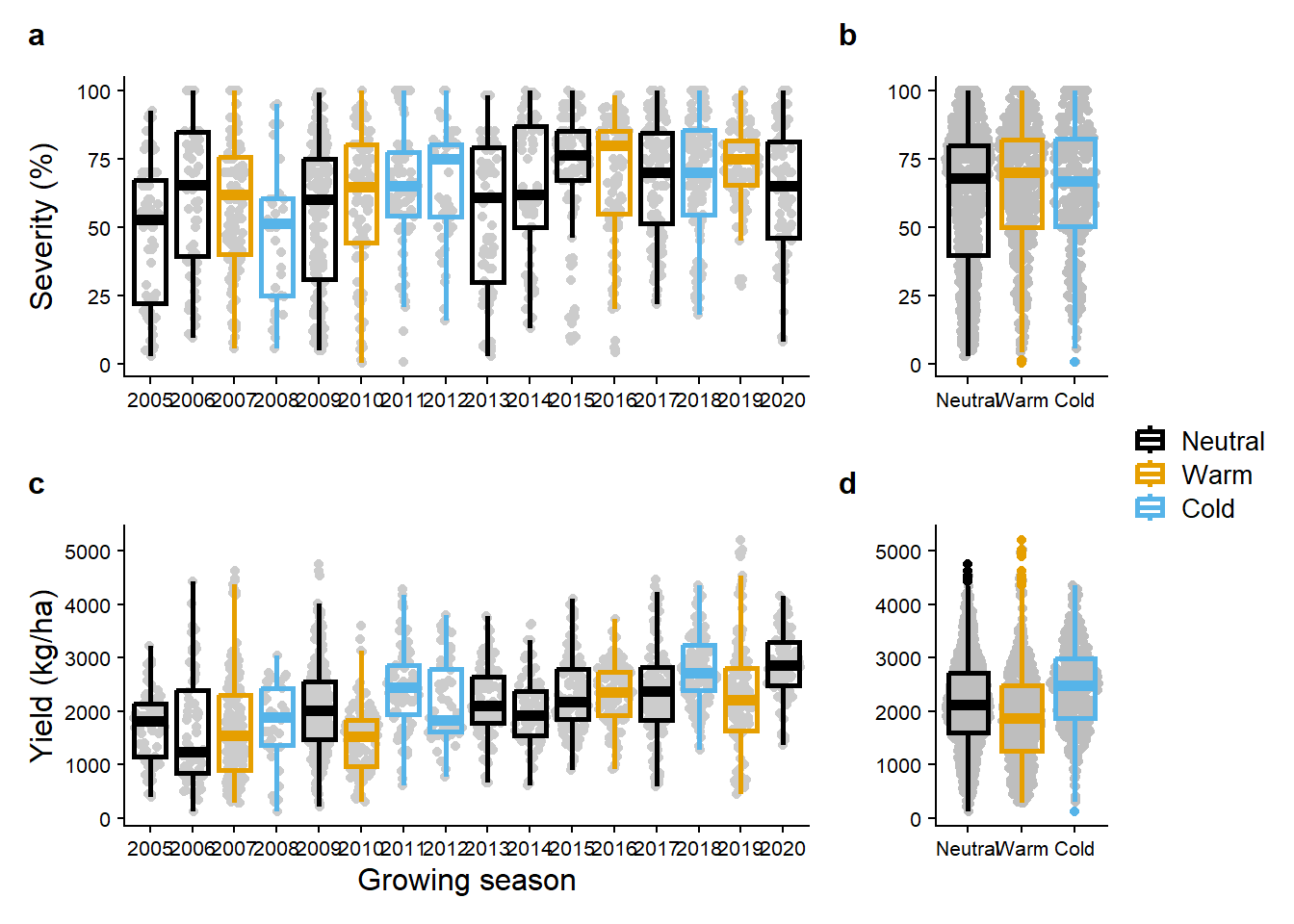
ggsave("figs/data_over_time.png", dpi = 600, height = 6, width = 9, bg = "white")
ggsave("figs/data_over_time.pdf", dpi = 600, height = 6, width = 9, bg = "white")Modeling disease damage
Yield vs. Severity
data_load2 |>
ggplot(aes(sev, yld, color = enso))+
# geom_point(alpha = 0.1)+
geom_smooth(aes(group=study),
method = "lm", size =0.1,se = F, fullrange =T)+
geom_smooth(aes(group = enso), color = "red", se = F, method = "lm")+
scale_color_colorblind()+
theme_half_open()+
facet_rep_grid(~enso)+
ylim(0,7500)+
labs(x = "Severity (%)",
y = "Yield (kg/ha)",
color = "")+
theme(strip.background = element_blank())`geom_smooth()` using formula = 'y ~ x'
`geom_smooth()` using formula = 'y ~ x'Warning: Removed 1124 rows containing missing values or values outside the scale range
(`geom_smooth()`).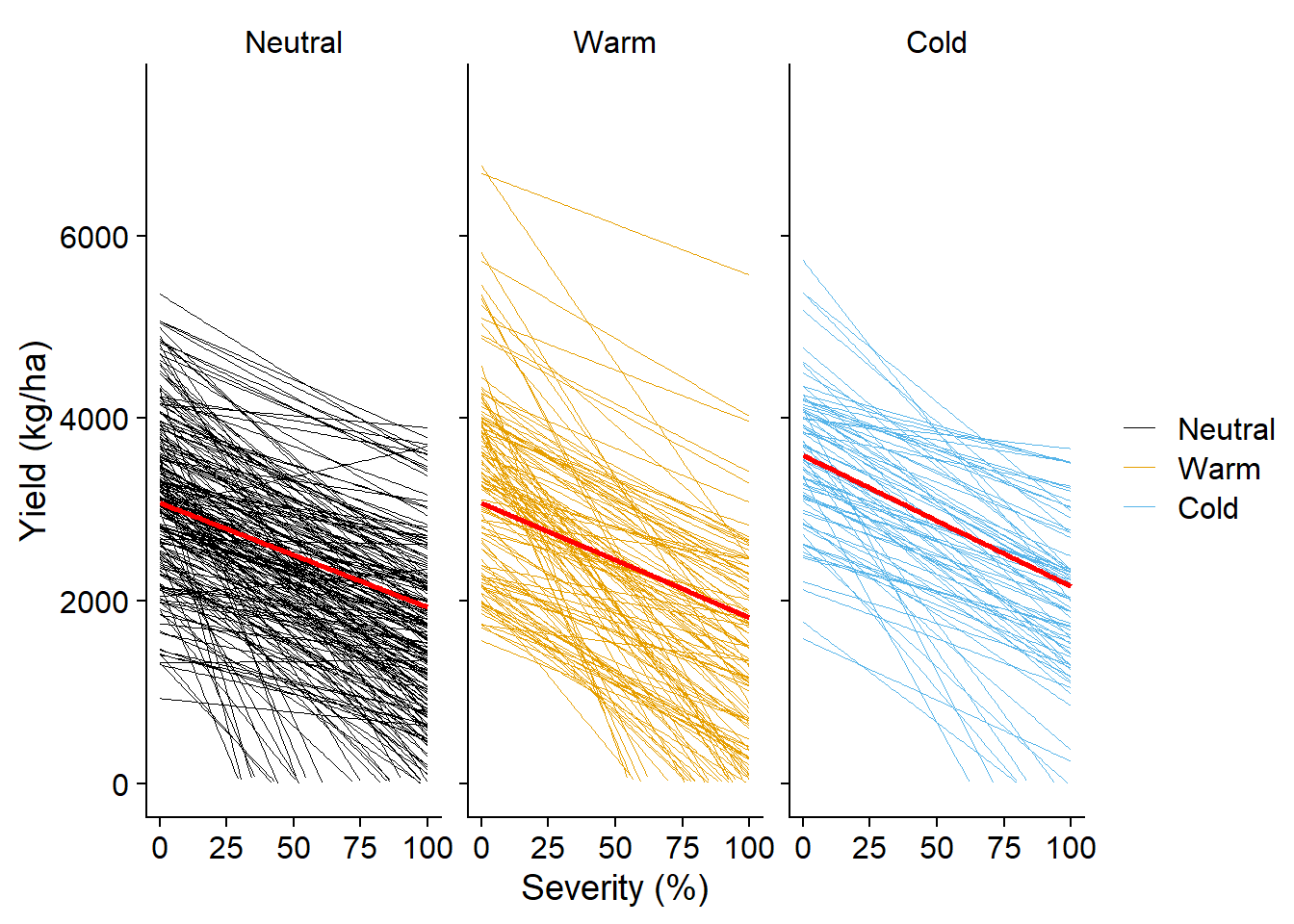
Meta-analysis
Ordinary Regression
reg_dc = data_load2 |>
group_by(study, year, region, enso) |>
summarise(intercept = lm(yld~sev)$coefficients[1],
slope = lm(yld~sev)$coefficients[2],
r2 = summary(lm(yld~sev))$r.squared,
sigma = summary(lm(yld~sev))$sigma) %>%
mutate(Dc = (slope/intercept)*100) |>
filter(Dc<0.5)`summarise()` has grouped output by 'study', 'year', 'region'. You can override
using the `.groups` argument.reg_dcGraph of the regression lines
reg1_gg = reg_dc |>
ggplot()+
geom_abline(aes(intercept = intercept, slope = slope, color= enso), alpha = 0.9)+
labs(x = "Severity (%)",
y = "Yield (kg/ha)")+
theme_half_open()+
xlim(0,100)+
ylim(0,7500)+
scale_color_colorblind()+
theme_half_open(font_size = 10)+
facet_rep_grid(~enso)+
labs(x = "SBR severity (%)",
y = "Soybean yield (kg/ha)",
color = "")+
theme(strip.background = element_blank(),
legend.position = "none")
reg1_gg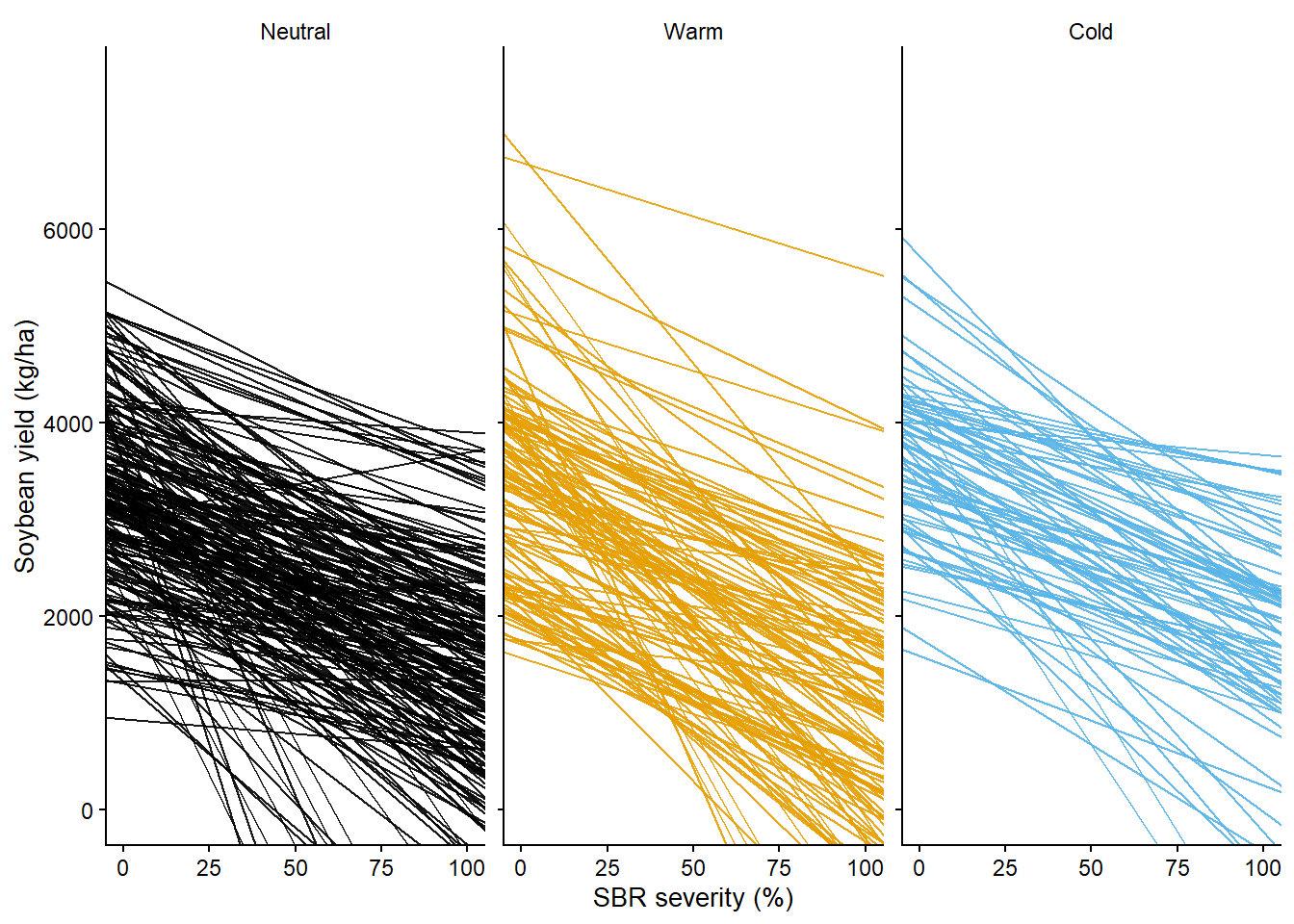
Distribution of intercepts and slopes
reg_dc |>
pivot_longer(c(intercept, slope), names_to = "par",values_to = "par_value")reg_dc |>
pivot_longer(c(intercept, slope), names_to = "par",values_to = "par_value") |>
group_by(par, enso) |>
summarise(avg = round(mean(par_value),1),
std_dev = round(sd(par_value),1),
q_lw = round(quantile(par_value,c(0.025)),1),
q_up = round(quantile(par_value,c(0.975)),1),
q_up-q_lw)`summarise()` has grouped output by 'par'. You can override using the `.groups`
argument.parms_gg = reg_dc |>
pivot_longer(c(intercept, slope), names_to = "par",values_to = "par_value") |>
mutate(par = ifelse(par=="intercept", "Intercept", "Slope")) |>
ggplot(aes(y = enso, par_value,color = enso))+
ggdist::stat_dotsinterval(slab_shape = 19,point_interval = "mean_qi", .width = c(0.95), quantiles = 500, slab_color= "gray60") +
scale_color_colorblind()+
theme_half_open(font_size = 10)+
labs(x = "Parameter estimate",
y = "")+
facet_rep_grid(~par, scales = "free")+
theme(strip.background = element_blank(),
legend.position = "none")
parms_gg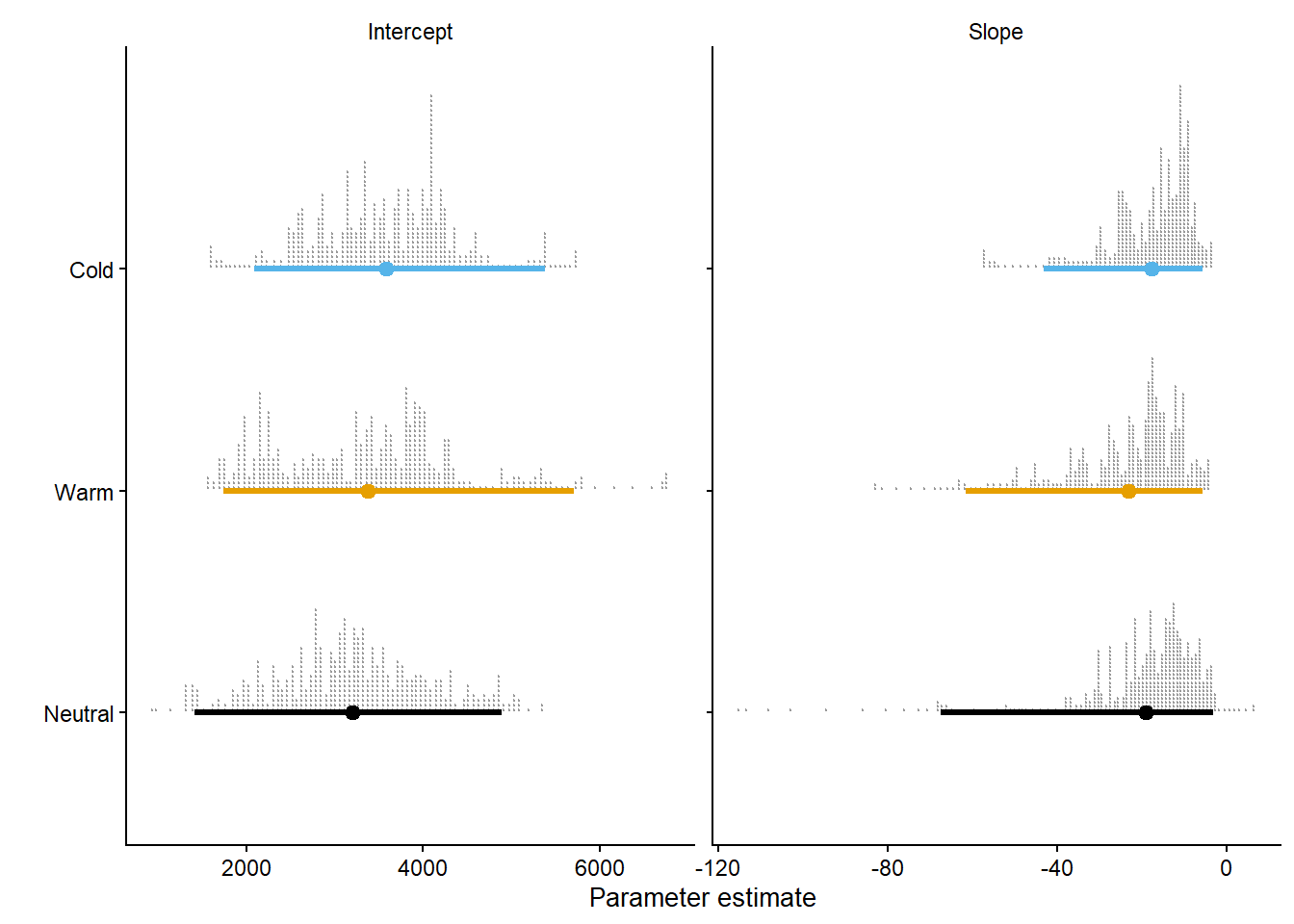
Relative yield loss (%) relative to β0
data2 = data_load2 |>
full_join(reg_dc) |>
mutate(l = 100*((intercept - yld)/intercept)) |>
filter(!is.na(l))Joining with `by = join_by(study, year, enso, region)`data2data2 |>
ggplot(aes(l))+
geom_histogram(color = "white", bins = 10)&
theme_half_open()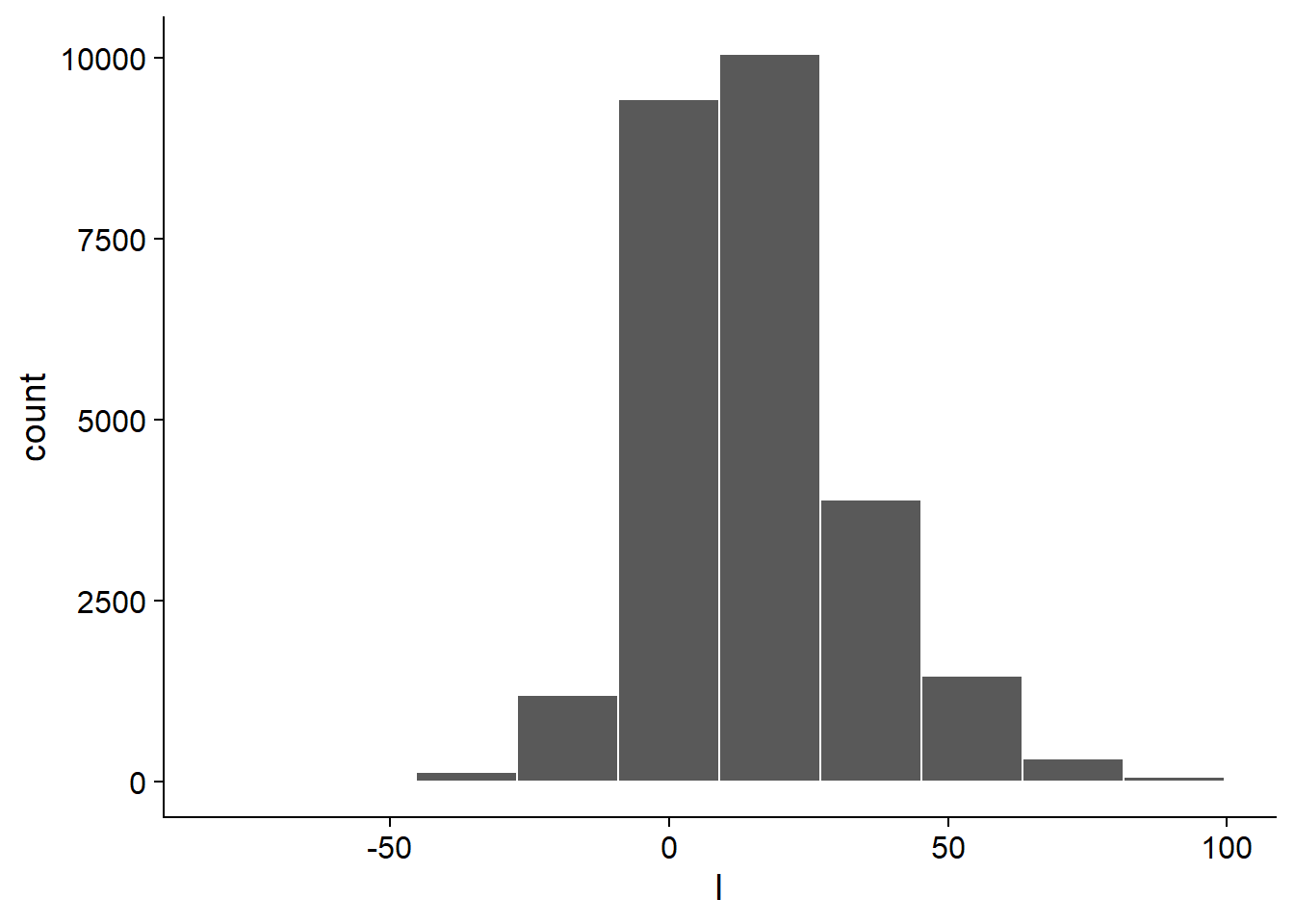
l_gg = data2 |>
ggplot(aes(sev,l,color =enso))+
geom_point( shape =1, alpha = 0.3)+
# color = "gray80",
# geom_smooth(method ="lm",
# aes(color =enso),
# color = "gray90",
# formula = y~0+x)+
geom_hline(yintercept = 0, linetype = "dashed", color = "gray40")+
scale_color_colorblind()+
labs(x = "Severity (%)",
y = expression("Yield loss relative to "~β[0]))+
theme_half_open(font_size = 10)+
facet_rep_grid(~enso)+
labs(x = "SBR severity (%)",
# y = expression("Yield loss relative to "~β[0]~"(%)"),
y = expression("Relative yield loss (%)"),
color = "")+
theme(strip.background = element_blank(),
legend.position = "none")
l_gg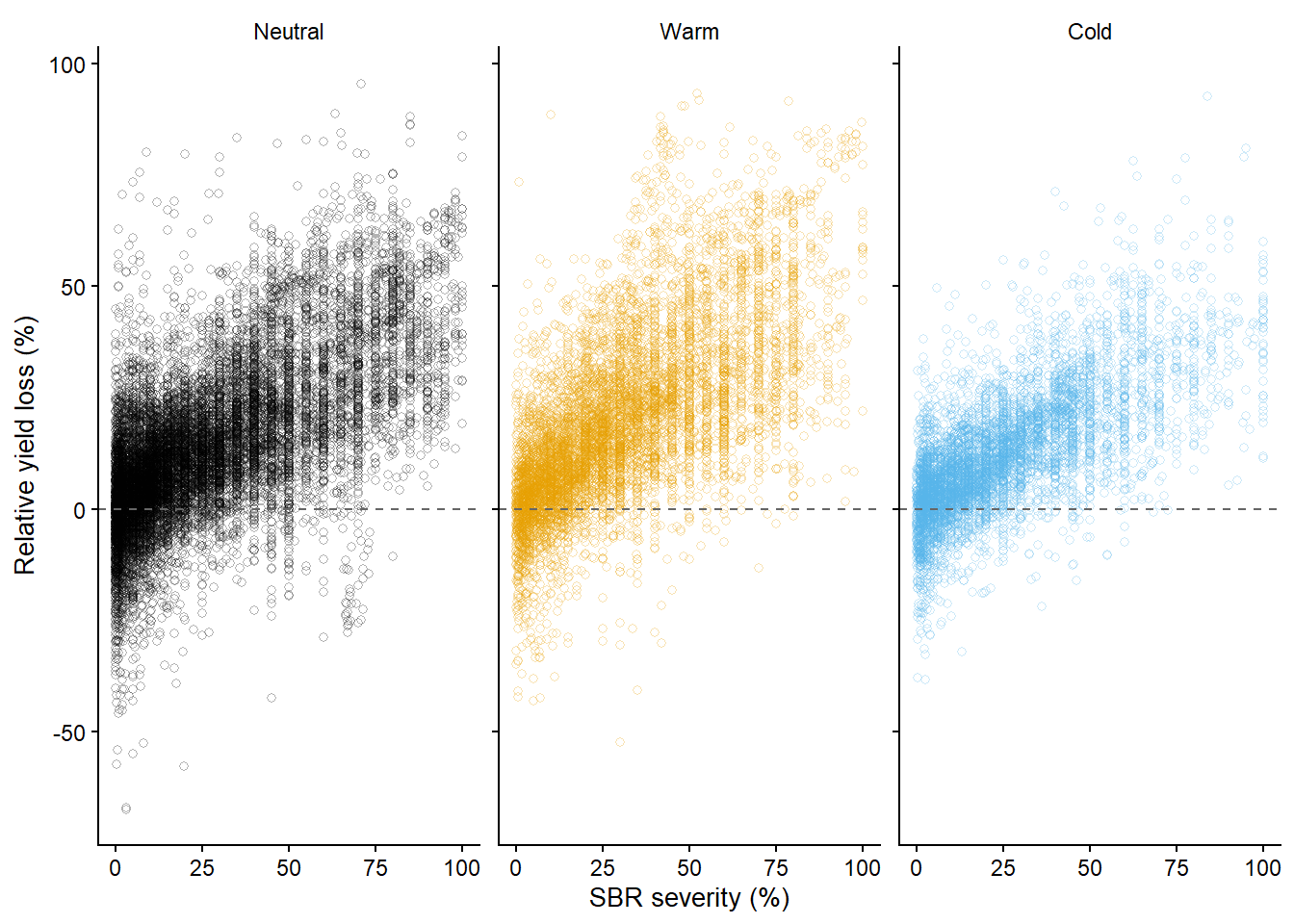
Second Regression l ~ 0 + sev
data3 = data2 |>
group_by(study, year, enso, region) |>
summarise(slope = lm(l~0+sev)$coefficients,
vi = as.numeric(vcov(lm(l~0+sev)))
) |>
full_join(enso_data_class)`summarise()` has grouped output by 'study', 'year', 'enso'. You can override
using the `.groups` argument.
Joining with `by = join_by(year, enso)`data3reg2_gg =
data3 |>
mutate(enso = factor(enso, levels = c("Neutral", "Warm", "Cold"))) |>
ggplot()+
geom_abline(aes(intercept = 0, slope = slope, color= enso), alpha = 0.9)+
labs(x = "Severity (%)",
y = "Yield (kg/ha)")+
theme_half_open()+
xlim(0,100)+
ylim(0,100)+
scale_x_continuous(expand = c(0, 0),
limits = c(0,100)) +
scale_y_continuous(expand = c(0, 0),
limits = c(0,100))+
scale_color_colorblind()+
theme_half_open(font_size = 10)+
facet_rep_grid(~enso)+
labs(x = "SBR severity (%)",
y = expression("Relative yield loss (%)"),
color = "")+
theme(strip.background = element_blank(),
legend.position = "none")Scale for x is already present.
Adding another scale for x, which will replace the existing scale.
Scale for y is already present.
Adding another scale for y, which will replace the existing scale.reg2_gg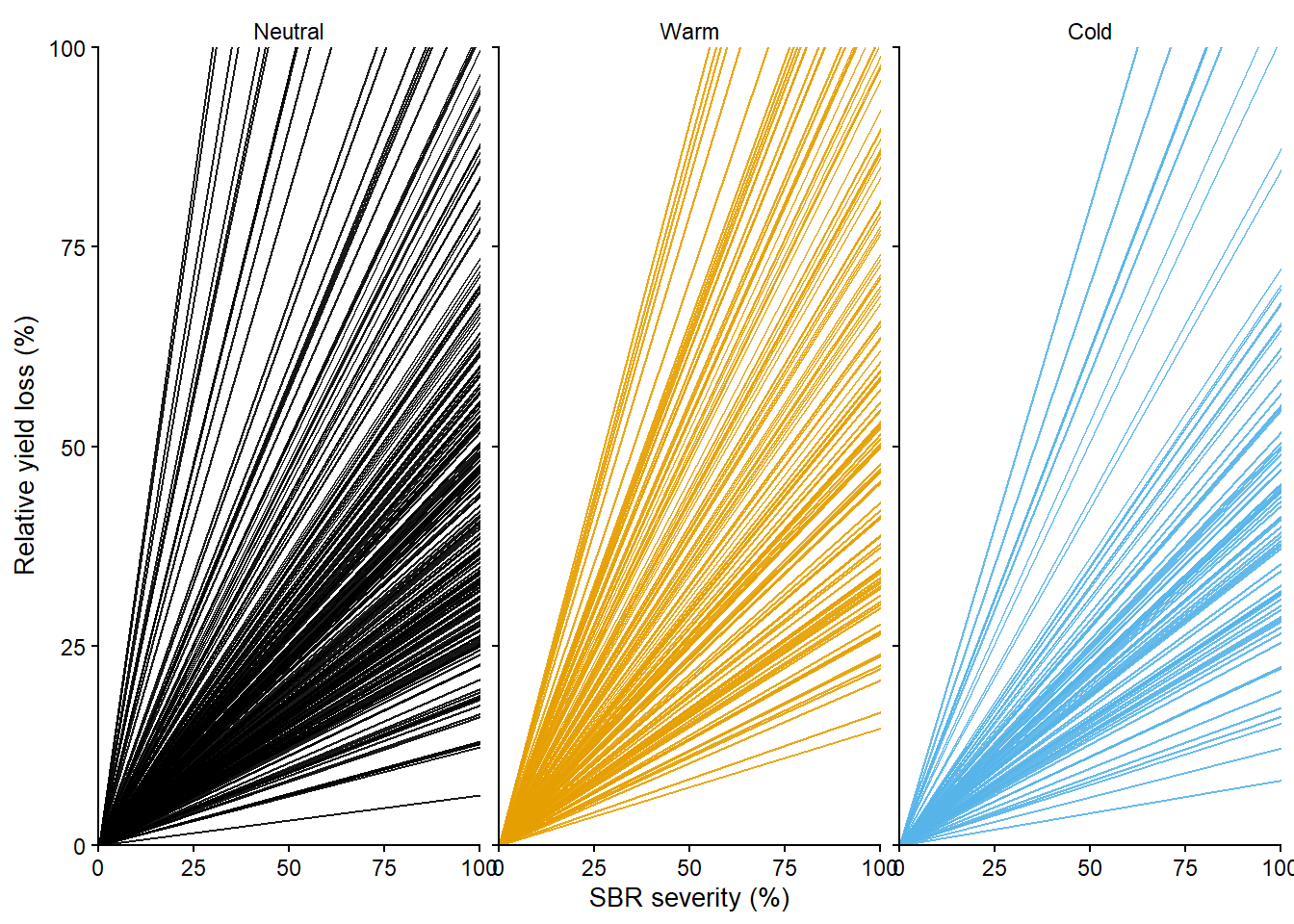
data3 |>
group_by(enso) |>
summarise(mean(slope),
median(slope),
round(quantile(vi,0.95),2))Damage vs. ENSO phases
data3 |>
ggplot(aes(slope))+
geom_histogram(color = "gray")+
facet_rep_wrap(~enso, ncol=1 )`stat_bin()` using `bins = 30`. Pick better value with `binwidth`.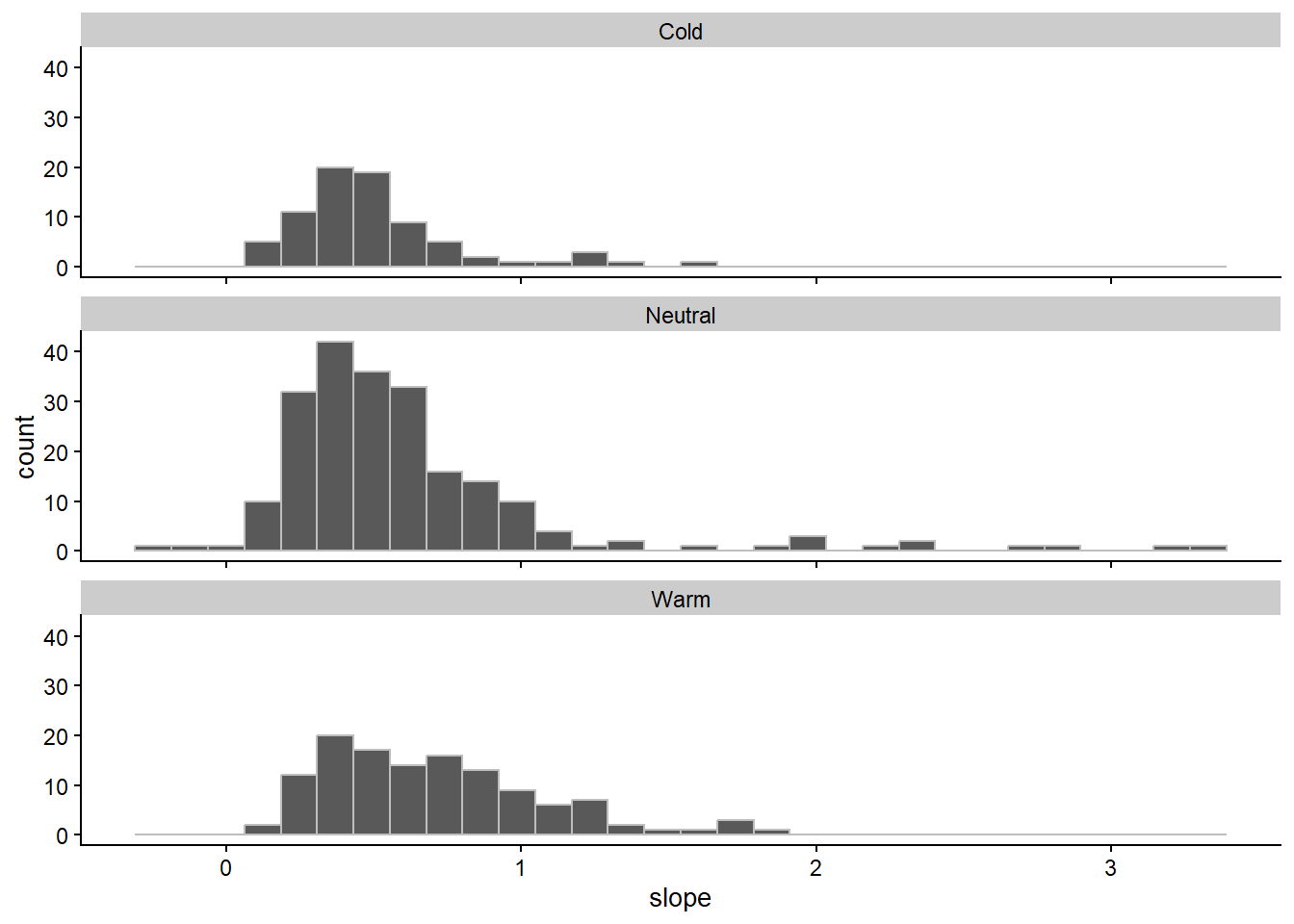
library(scales)
Anexando pacote: 'scales'O seguinte objeto é mascarado por 'package:purrr':
discardO seguinte objeto é mascarado por 'package:readr':
col_factorslopes_gg = data3 |>
mutate(enso = factor(enso, levels = c("Neutral", "Warm", "Cold"))) |>
arrange(slope) |>
ggplot(aes(enso, slope ))+
geom_boxplot(fill = NA, color = "steelblue", size =1, outlier.colour = NA)+
geom_sina(aes(size = vi, color= vi), alpha = 0.5)+
scale_size_continuous(range = c(1, 7),
#limits = c(0.01,1),
breaks = c(0.001, 0.01, 0.1, 1),
label=scientific_format())+
scale_color_gradient(low = "black",
high = "red",
#limits = c(0.01, 1),
breaks = c(0.001, 0.01, 0.1, 1),
label=scientific_format())+
guides(color= guide_legend(),
size=guide_legend())+
theme_half_open(font_size = 10)+
labs(y = "Study-level damage coefficient (%/pp)",
x = "",
size = "Slope variance",
color = "Slope variance" )+
theme(legend.position = "top")
slopes_gg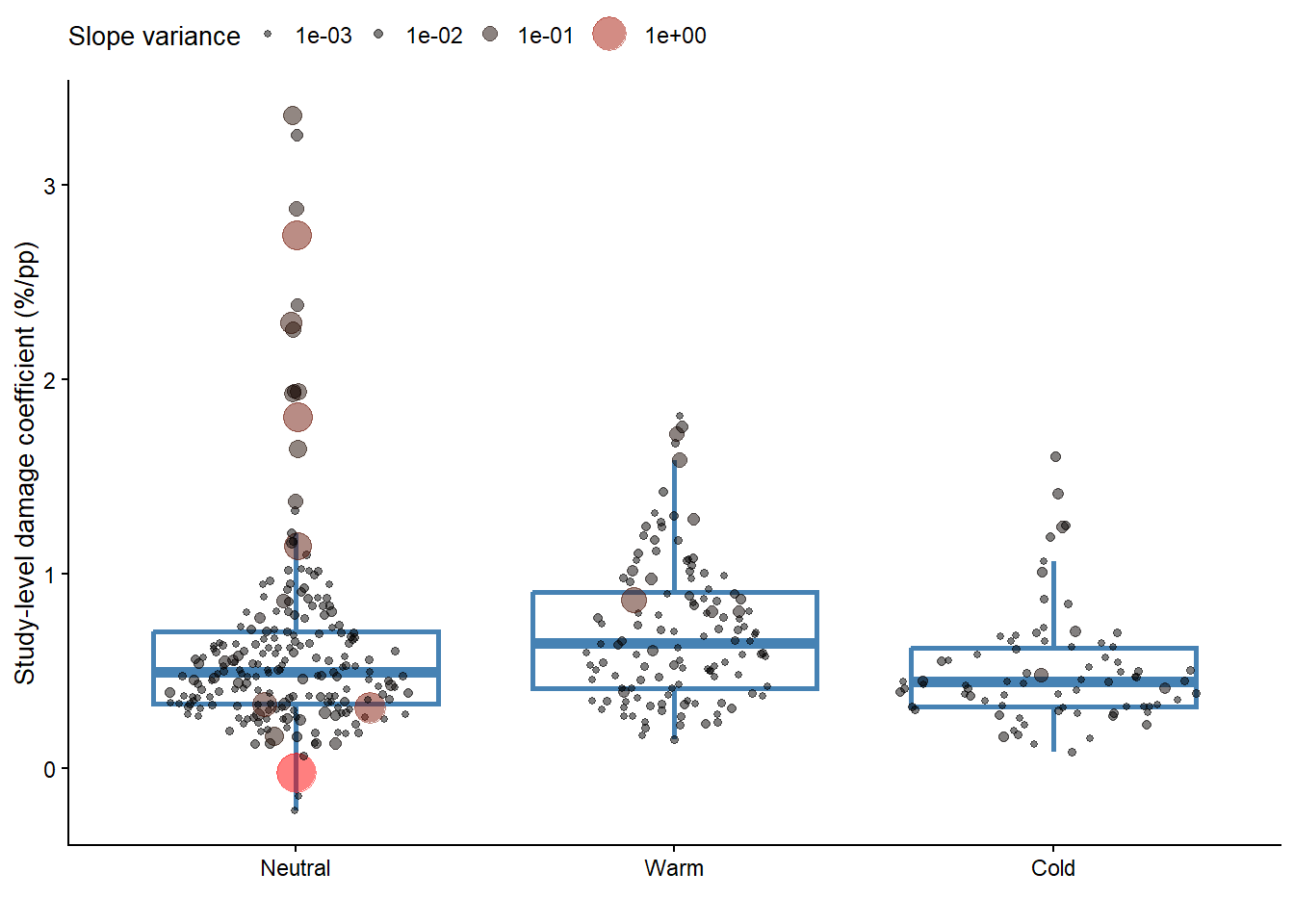
Combo plot
layout <- "
AAAACC
BBBBCC
"
layout <- "
AAAADD
BBBBDD
CCCCDD
"
layout <- "
AAAABBB
CCCCEEE
DDDDEEE
"
layout <- "
AAAABBB
AAAABBB
AAAABBB
CCCCBBB
CCCCEEE
DDDDEEE
DDDDEEE
DDDDEEE
"
# (reg1_gg+parms_gg+l_gg+reg2_gg+slopes_gg)+
((reg1_gg/l_gg/reg2_gg) | (parms_gg/slopes_gg))+
plot_annotation(tag_levels = 'a')&
theme(axis.title = element_text(size = 10)
# plot.tag = element_text(size = 8)
)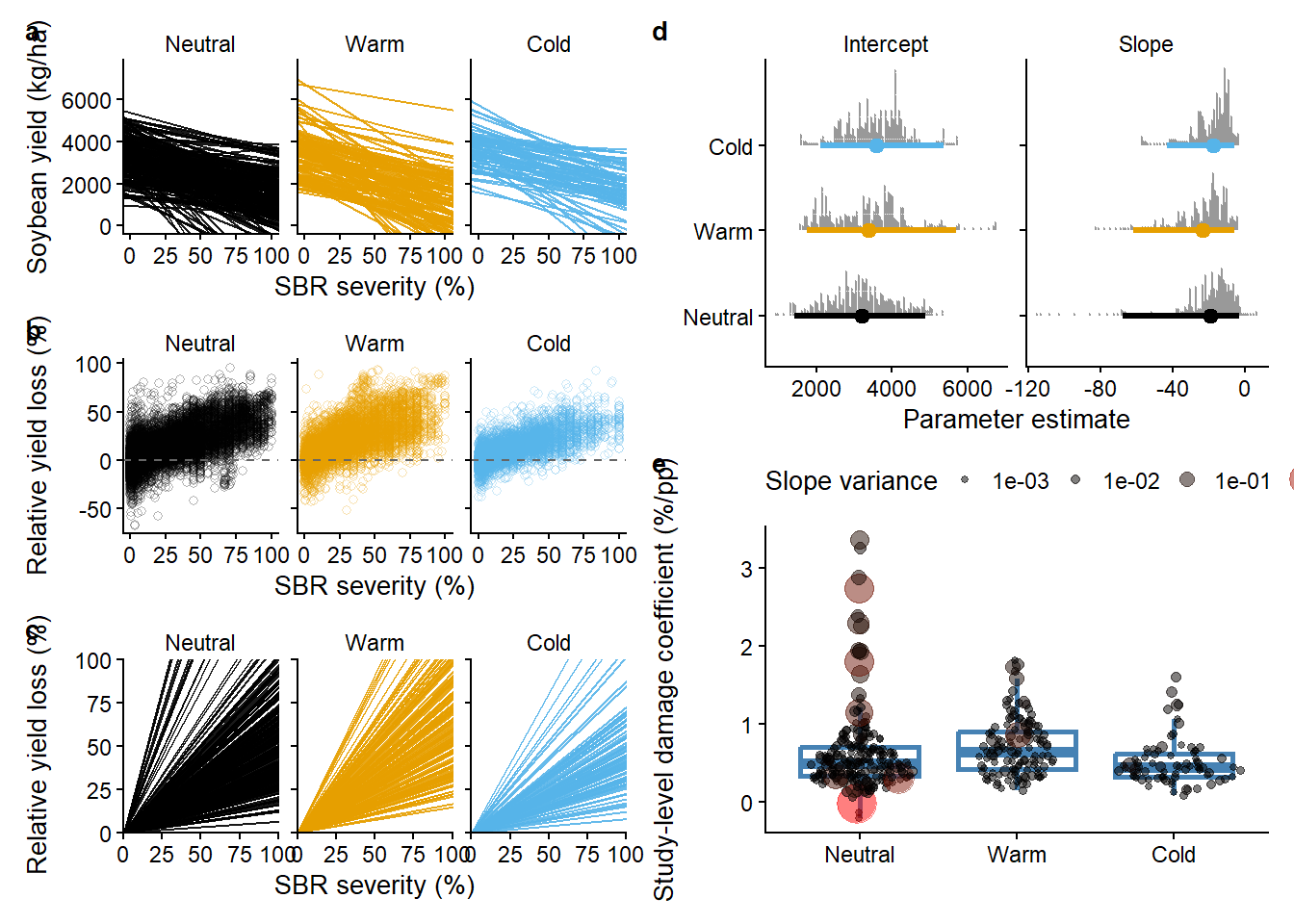
ggsave("figs/regression_lines.png", dpi = 600, width = 10, height = 6, bg ="white")
ggsave("figs/regression_lines.pdf", dpi = 600, width = 10, height = 6, bg ="white")Meta-analytical model
data4 = data3 |>
mutate(year = as.factor(year),
enso = factor(enso, levels = c("Neutral", "Cold", "Warm")))
metamodel2 = rma.uni(yi = slope,
vi = vi,
mods = ~ enso,
random = list(~1|year/study),
# struct = "HCS",
method = "ML",
data =data4)Warning: Extra argument ('random') disregarded.metamodel2
Mixed-Effects Model (k = 417; tau^2 estimator: ML)
tau^2 (estimated amount of residual heterogeneity): 0.1099 (SE = 0.0080)
tau (square root of estimated tau^2 value): 0.3316
I^2 (residual heterogeneity / unaccounted variability): 99.54%
H^2 (unaccounted variability / sampling variability): 219.29
R^2 (amount of heterogeneity accounted for): 5.61%
Test for Residual Heterogeneity:
QE(df = 414) = 41096.3735, p-val < .0001
Test of Moderators (coefficients 2:3):
QM(df = 2) = 17.5026, p-val = 0.0002
Model Results:
estimate se zval pval ci.lb ci.ub
intrcpt 0.5651 0.0236 23.9141 <.0001 0.5188 0.6114 ***
ensoCold -0.0640 0.0449 -1.4237 0.1545 -0.1520 0.0241
ensoWarm 0.1258 0.0385 3.2649 0.0011 0.0503 0.2014 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1data4 = data3 |> mutate(year = as.factor(year))
metamodel2 = rma.uni(yi = slope,
vi = vi,
mods = ~ 0+enso,
random = list(~1|year/study),
# struct = "HCS",
method = "ML",
data =data4)Warning: Extra argument ('random') disregarded.metamodel2
Mixed-Effects Model (k = 417; tau^2 estimator: ML)
tau^2 (estimated amount of residual heterogeneity): 0.1099 (SE = 0.0080)
tau (square root of estimated tau^2 value): 0.3316
I^2 (residual heterogeneity / unaccounted variability): 99.54%
H^2 (unaccounted variability / sampling variability): 219.29
Test for Residual Heterogeneity:
QE(df = 414) = 41096.3735, p-val < .0001
Test of Moderators (coefficients 1:3):
QM(df = 3) = 1258.6134, p-val < .0001
Model Results:
estimate se zval pval ci.lb ci.ub
ensoCold 0.5011 0.0382 13.1116 <.0001 0.4262 0.5760 ***
ensoNeutral 0.5651 0.0236 23.9141 <.0001 0.5188 0.6114 ***
ensoWarm 0.6909 0.0305 22.6896 <.0001 0.6312 0.7506 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1Estimates for each moderator
grid = qdrg(object = metamodel2, data = data4, at = list(vi = 0, year.c = 0))
# cld(emmeans(grid,specs = ~enso, by = "region"), Letters = letters)
cld(emmeans(grid, specs = ~enso), Letters = letters)# grid@VGraphs
Forest plot
forest_gg = data3 |>
mutate(study = as.character(study),
sd = sqrt(vi),
cil = slope - 1.96*sd,
ciu = slope + 1.96*sd) |>
full_join(as.data.frame(emmeans(grid,specs = ~enso))) |>
mutate(enso = factor(enso, levels = c("Neutral", "Warm","Cold"))) |>
ggplot(aes(slope,reorder(study, slope), color= enso))+
geom_point(size=0.3)+
geom_errorbar(aes(xmin=cil, xmax = ciu), width = 0,size=0.3, alpha =0.5)+
geom_vline(xintercept = 0, linetype = "dashed")+
geom_vline(aes(xintercept = emmean), color = "gray40", size = .5)+
scale_color_colorblind()+
facet_rep_wrap(~enso, scales="free_y")+
labs(y = "",
x = "Damage coefficient (%/p.p.)",
color = "")+
guides(color = guide_legend(override.aes = list(size=2.5)))+
theme(axis.text.y = element_blank(),
axis.ticks.length.y = unit(0, "cm"),
strip.background = element_blank(),
legend.position = "bottom")Joining with `by = join_by(enso)`forest_gg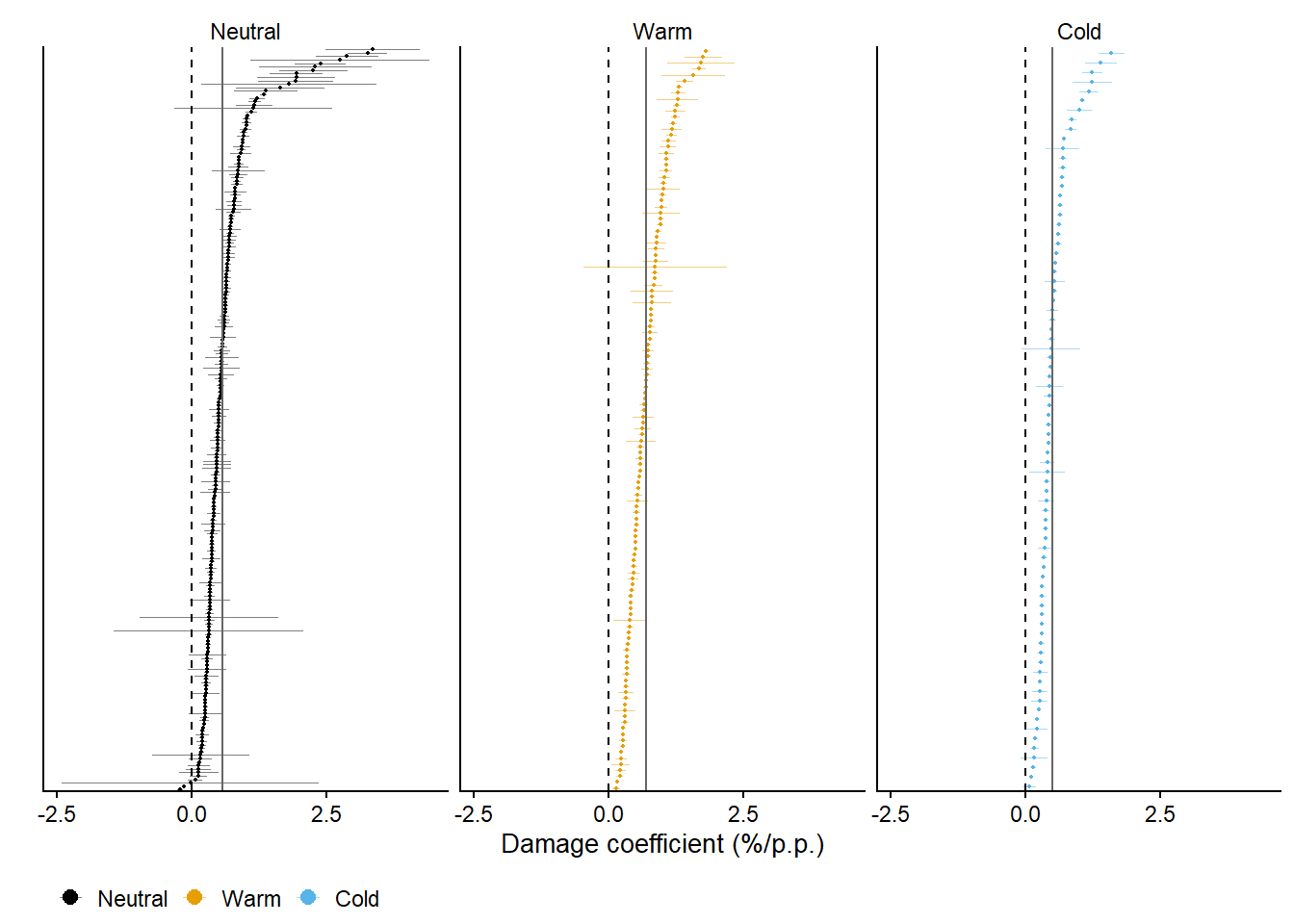
Damage coefficient
dc_data = as.data.frame(emmeans(grid, specs = ~enso))
dc_dataDC_gg = as.data.frame(emmeans(grid,specs = ~enso)) |>
mutate(enso = factor(enso, levels = c("Neutral", "Warm","Cold"))) |>
ggplot(aes(reorder(enso,emmean), emmean, color = enso))+
geom_point(position = position_dodge(width = 0.2), size= 3)+
geom_errorbar(aes(ymin =lower.CL, ymax = upper.CL),
position = position_dodge(width = 0.2),
width = 0,
size = 0.7)+
labs(x = "",
y = "Damage coefficient (%/p.p.)",
color = "")+
scale_y_continuous(breaks = seq(0.1,0.8, by = 0.1), limits =c(0.35,0.81))+
scale_color_colorblind()+
theme(legend.position = "none")
DC_gg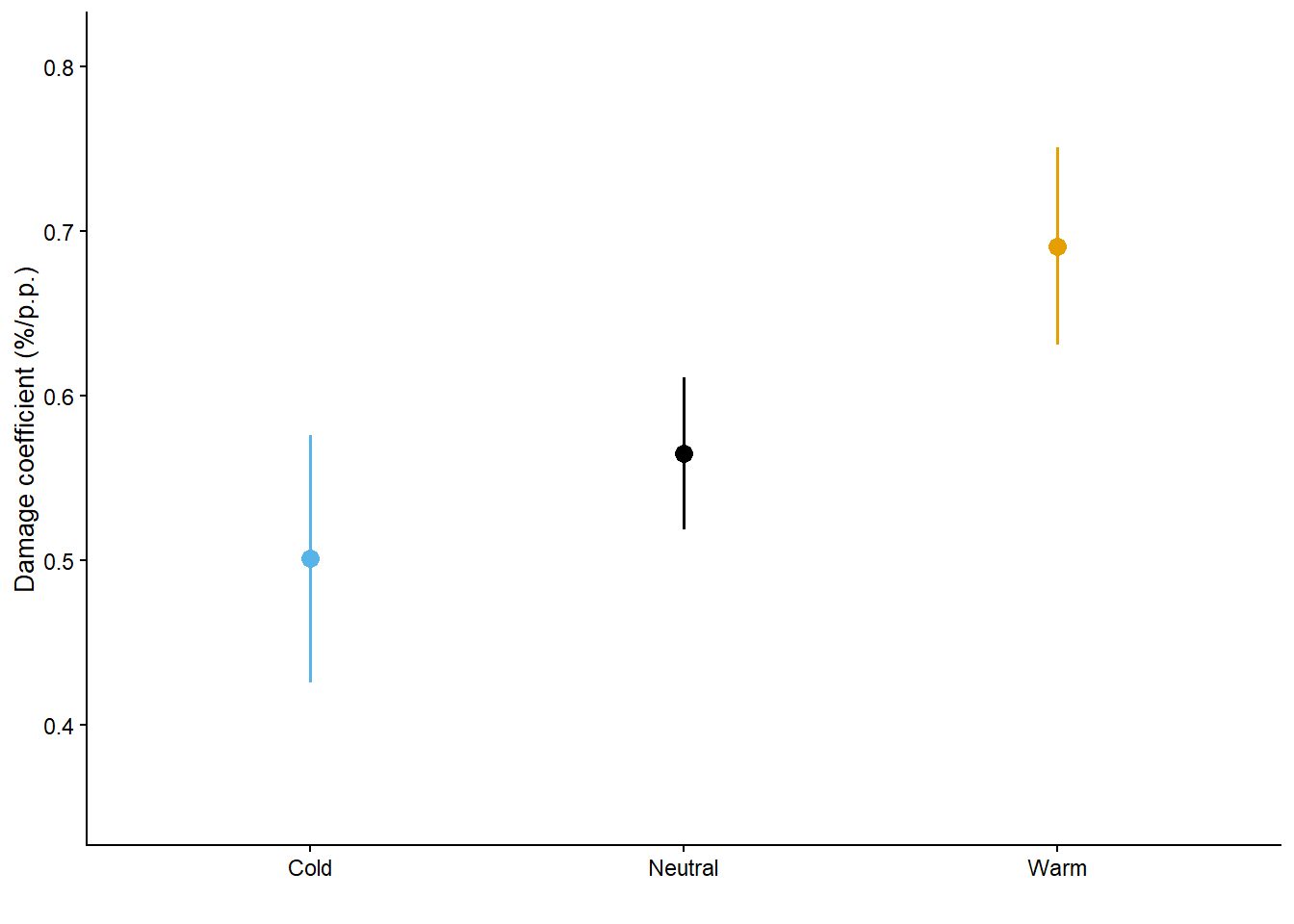
Yield loss
damage_coef_df =as.data.frame(emmeans(grid,specs = ~enso)) |>
mutate(enso = factor(enso, levels = c("Neutral", "Warm","Cold"))) |>
mutate(`100` = 100-100*emmean,
`0` = 100,
`50` = 100-50*emmean,
`50_upper` = 100-50*lower.CL,
`50_lower` = 100-50*upper.CL) |>
mutate(yield_50 = 100-50*emmean)
damage_coef_df_for_plot = damage_coef_df|>
pivot_longer(7:8,
names_to = "sev",
values_to = "yloss") %>%
mutate(sev = as.numeric(sev)) %>%
mutate(cil = 100-sev*lower.CL,
ciu = 100-sev*upper.CL) |>
mutate(yl = -(yloss-100))
rel_gg = damage_coef_df_for_plot |>
ggplot()+
geom_ribbon(aes(x= sev,ymin = cil, ymax = ciu, fill = enso ),alpha = 0.5, color = NA)+
geom_line(aes(sev, yloss, color = enso),size = 1)+
geom_vline(xintercept =50, linetype = "dashed", color = "gray40", size = 0.2)+
geom_hline(aes(yintercept = yield_50, color = enso), linetype = "dashed", size = 0.2)+
geom_point(aes(x = 50, y = yield_50, fill =enso ), size = 2, shape = 21, color = "white")+
geom_label_repel(data = damage_coef_df,
aes(x =50, y = yield_50, fill =enso, label = paste0("Yield = ", round(yield_50,1),"%")),
label.size = 0.1,
size = 2,
color = "white",
seed = 123,
show.legend = F)+
guides(text = F)+
scale_linetype_manual(values = 2)+
scale_color_colorblind()+
scale_fill_colorblind()+
scale_x_continuous(limits = c(0,100))+
scale_y_continuous(limits = c(0,100), breaks = c(seq(0,100,by = 20)),
)+
theme(strip.background = element_blank(),
legend.position = "top")+
coord_equal()+
labs(x = "SBR Severity (%)",
y = "Soybean yield (%)",
color = "",
fill = "")Warning: The `<scale>` argument of `guides()` cannot be `FALSE`. Use "none" instead as
of ggplot2 3.3.4.rel_gg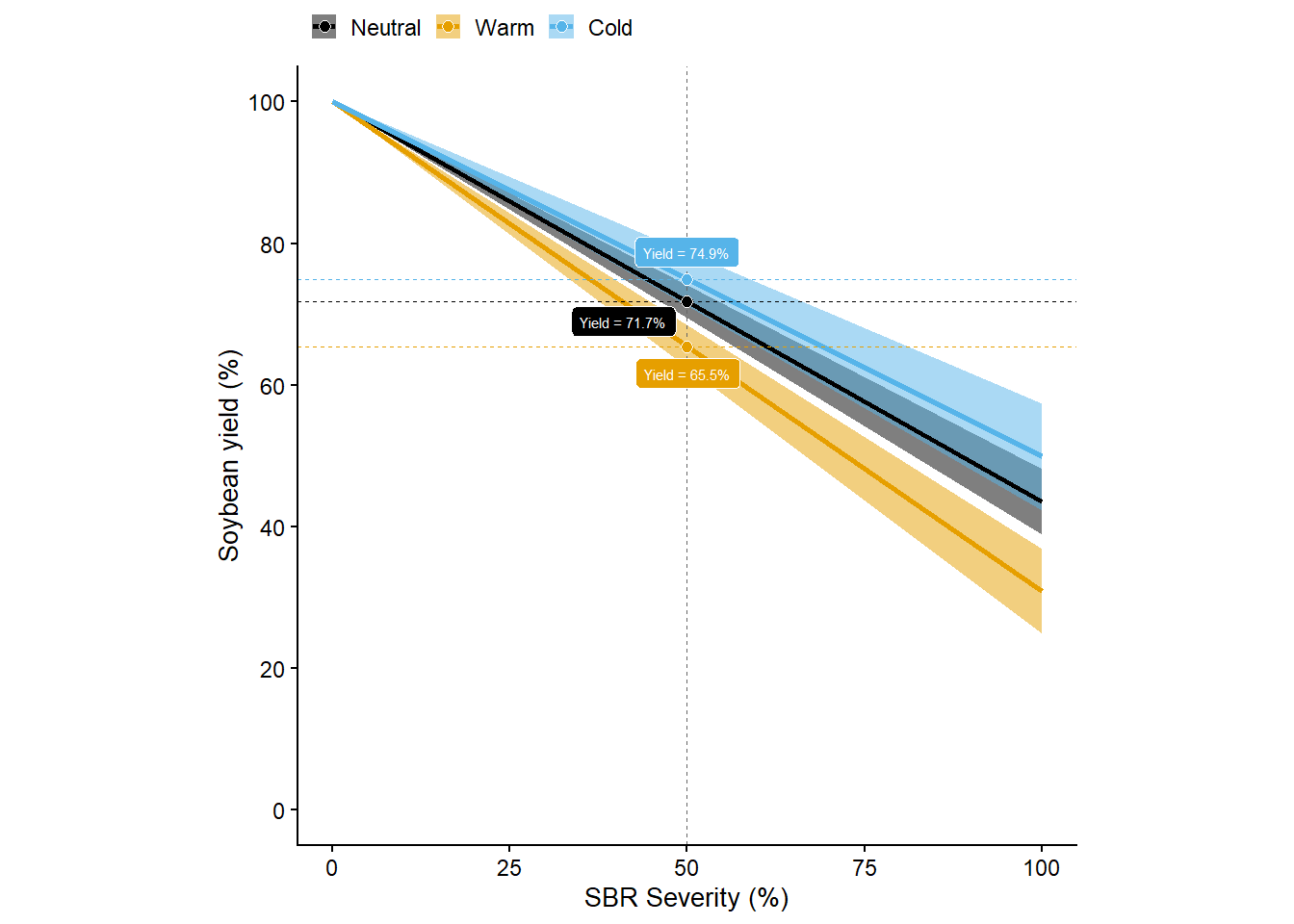
Combo plot
(DC_gg+rel_gg)+
plot_annotation(tag_levels = "a")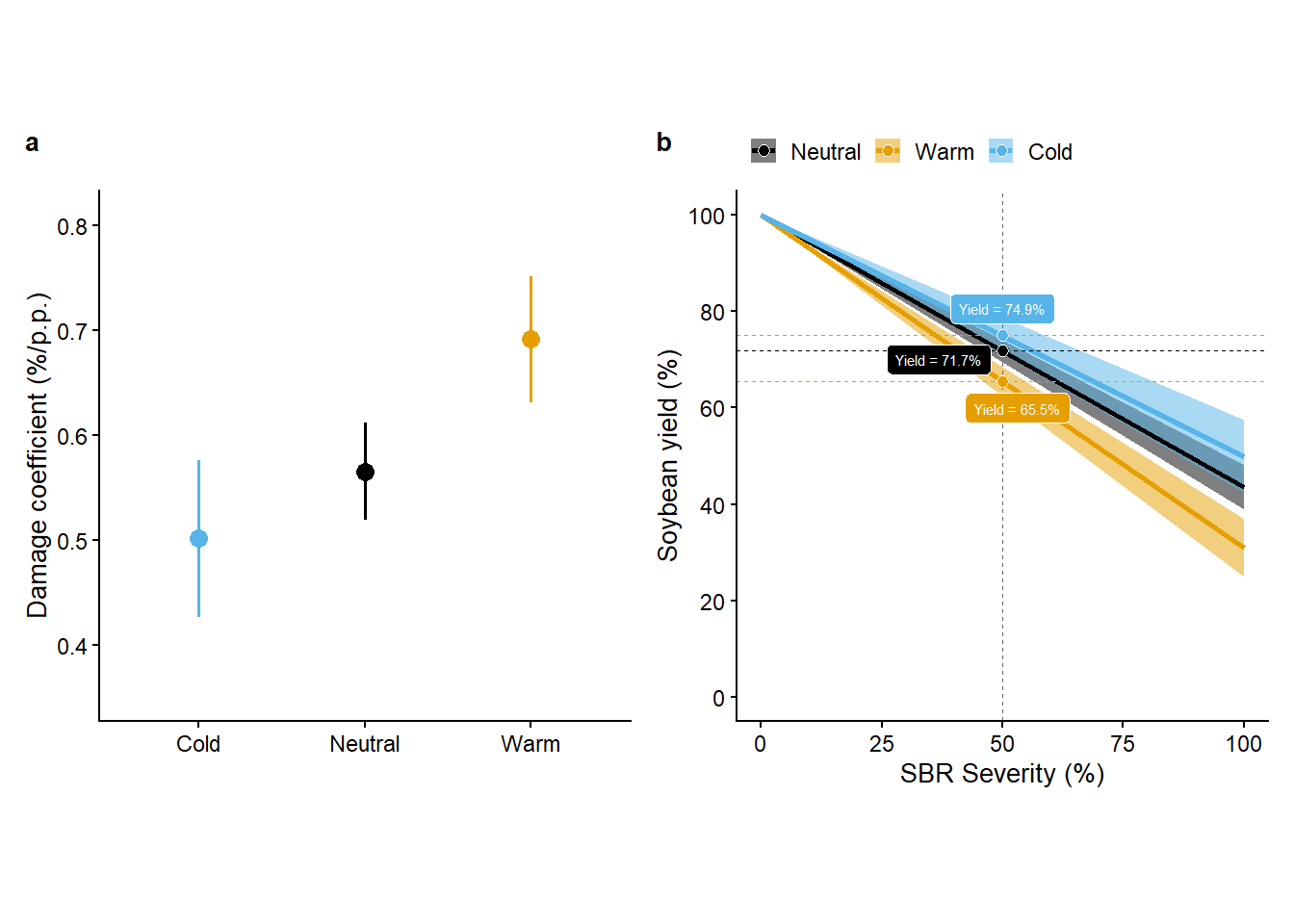
#
# plot_layout(guides = "collect") &
# theme(axis.title = element_text(size = 5),
# legend.position = "bottom")
ggsave("figs/z.png", dpi = 900, width =6, height = 3.5, bg ="white")
ggsave("figs/z.pdf", dpi = 900, width =6, height = 3.5, bg ="white")Yield protection
Vector for attainable yield (
ymax), control efficacy (lambda), severity in untreated field (sn), and damage coefficient (a).
length_grid = 500
ymax = seq(1500, 4000,length.out = length_grid) # attainable yield
lambda = c(30, 50, 70) # control efficacy
sn = seq(0, 100,length.out = length_grid) # seveiry on untreated
a = dc_data$emmean # damage coeficientsCalculating yield protection (
yld_protection) for the grid of vectors
yprotection_df = expand.grid(ymax = ymax,lambda = lambda, sn = sn, a = a) |>
mutate(yld_protection = ((a*ymax)/100) *(sn-(sn*(1-lambda/100)))) |>
mutate(lambda = paste0(lambda,"% of Control")) |>
full_join(dc_data |> rename(a = emmean))Joining with `by = join_by(a)`Response surfaces
Absolute yield protection
yprotection_df |>
ggplot(aes(sn, ymax, fill = yld_protection))+
geom_raster()+
scale_fill_viridis_b(option = "A",
guide = guide_colorbar(barwidth = 15, barheight = 0.3),
breaks = seq(0, 3000, by =250)
)+
facet_grid(lambda~enso)+
scale_y_continuous(breaks = seq(min(ymax), max(ymax), length.out = 5))+
theme_minimal_grid(font_size = 10)+
labs(y = "Attainable yield (kg/ha)",
x = "Severity untreated (%)",
fill ="Yield protection (kg/ha)" )+
theme(panel.grid = element_blank(),
axis.text = element_text(size = 5),
legend.position = "bottom")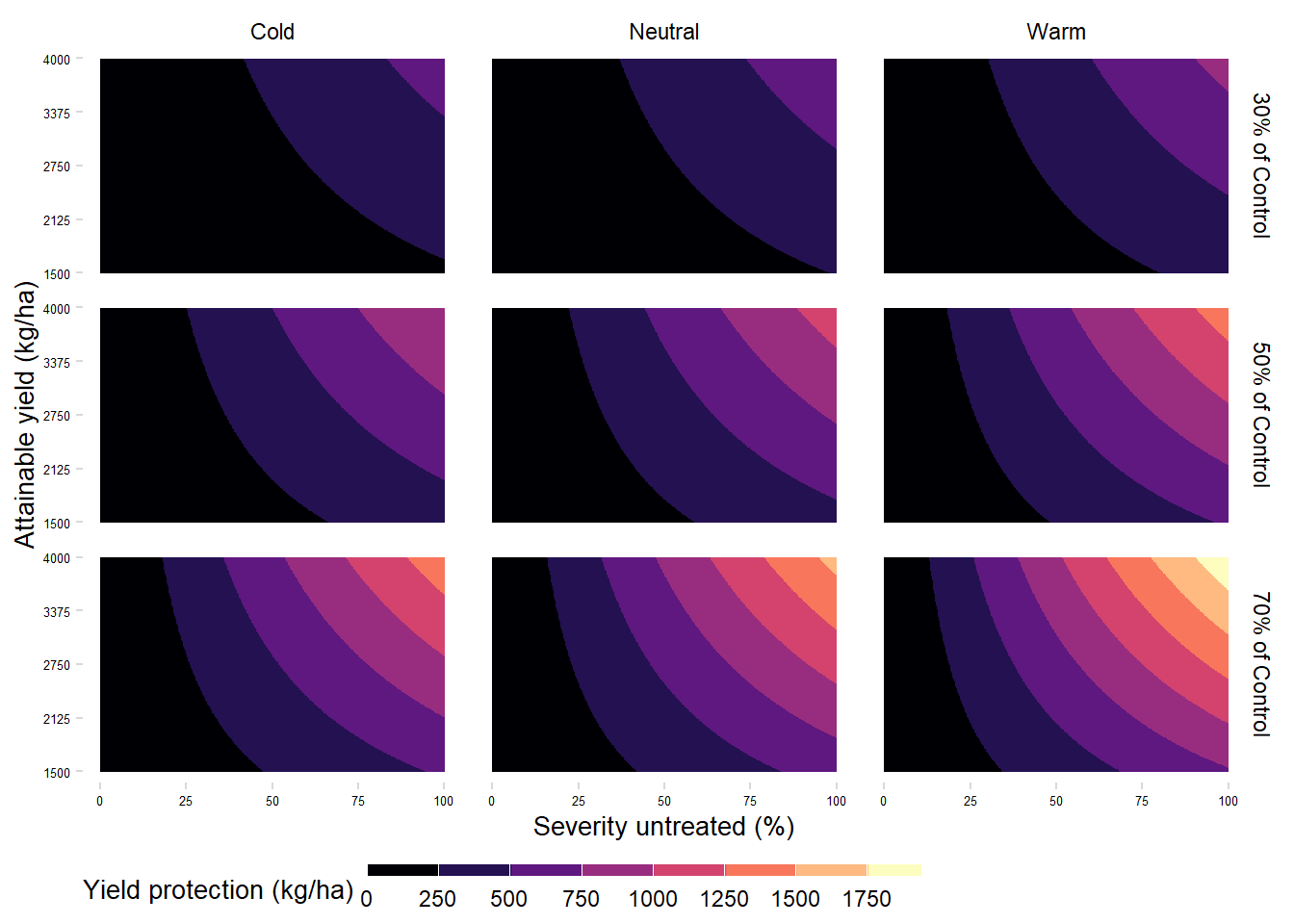
ggsave("figs/surface_yield_protection.png", dpi = 600,height =5, width = 5, bg = "white" )Yield protection difference from Neutral ENSO
yprotection_df_diff = yprotection_df |>
group_by(enso) |>
mutate(id = 1:length(enso)) |>
ungroup() |>
pivot_wider(id_cols = c(id,lambda, sn,ymax),
values_from = yld_protection,
names_from = enso) |>
mutate(Cold = Cold - Neutral,
Warm = Warm -Neutral) |>
dplyr::select(-Neutral) |>
pivot_longer(5:6,
values_to = "diff",
names_to = "enso")
max(yprotection_df_diff$diff)[1] 352.3533yprotection_df_diff |>
ggplot(aes(sn, ymax, fill = diff))+
geom_raster()+
scale_fill_steps2(midpoint = 0,
low = "#742881ff",#"#",
mid = "#f9f9f9ff",
high = "#1b7939ff",#"#27456e",
guide = guide_colorbar(barwidth = 10, barheight = 0.3),
limits = c(min(yprotection_df_diff$diff),max(yprotection_df_diff$diff)),
breaks = seq(-200, 350, by =50))+
scale_y_continuous(breaks = seq(min(ymax), max(ymax), length.out = 5))+
scale_x_continuous(breaks = seq(min(sn)*100, max(sn)*100, length.out = 5 ))+
facet_grid(lambda~enso)+
theme_minimal_grid(font_size = 10)+
labs(y = "Attainable yield (kg/ha)",
x = "Severity untreated (%)",
fill ="Difference in yield protection\nfrom neutral ENSO phase (kg/ha)" )+
theme(panel.grid = element_blank(),
legend.position = "bottom",
axis.text = element_text(size =8),
legend.text = element_text(size =6),
legend.title = element_text(size =6))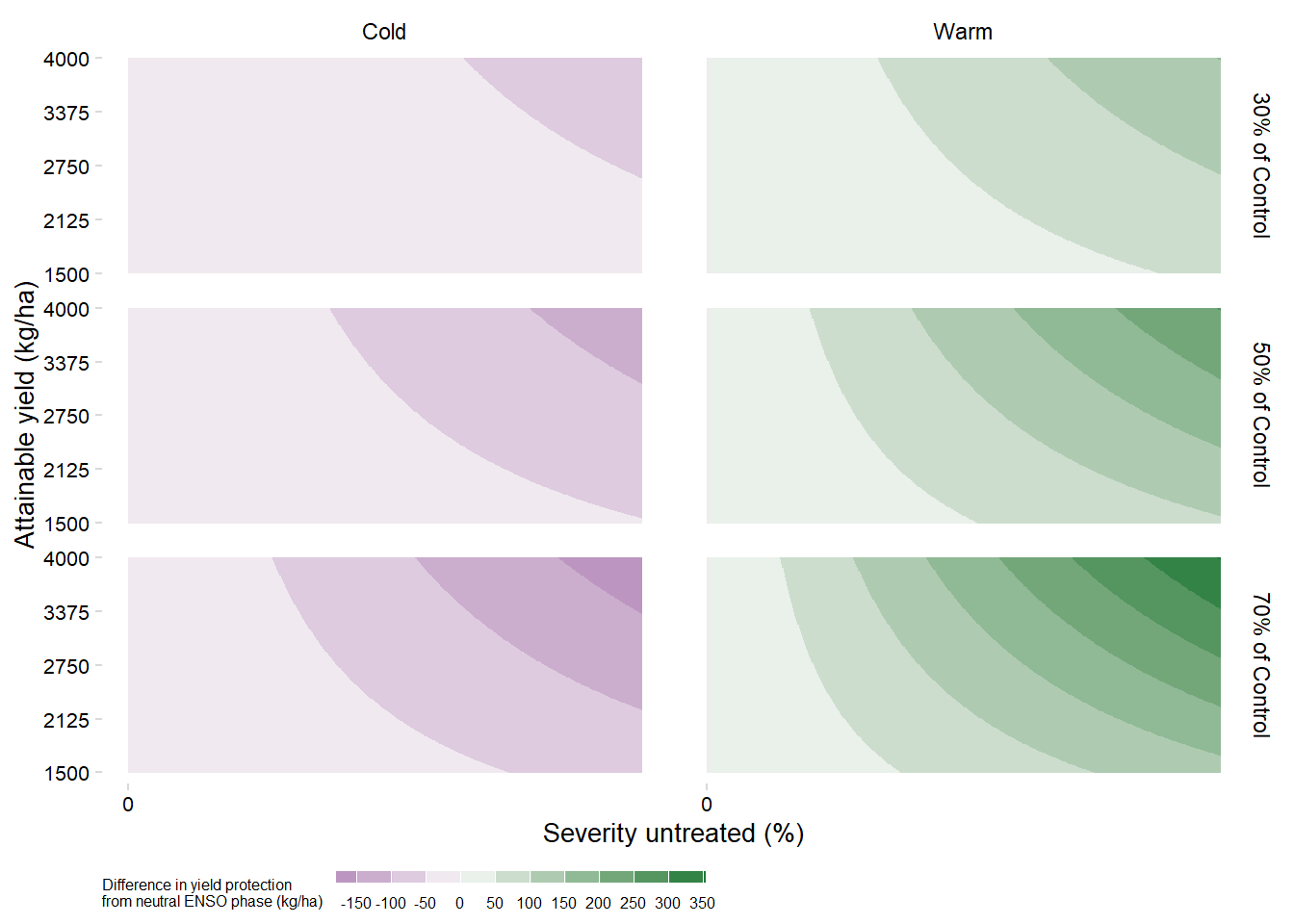
ggsave("figs/surface_yield_protection_difference.png", dpi = 600, height =5, width = 4, bg = "white" )
ggsave("figs/surface_yield_protection_difference.pdf", dpi = 600, height =5, width = 4, bg = "white" )expand.grid(a = seq(0.1, 1, length.out = 20),
sev = seq(0,100,by =5 ),
lambda = c(50, 70),
ymax = ymax[250])|>
mutate(yld_protection = (a*ymax*(sev - sev*((1-lambda/100))))/100) |>
ggplot(aes(sev, yld_protection, group = a, color = a))+
geom_line(size = 1)+
scale_color_viridis_c()+
facet_rep_wrap(~lambda)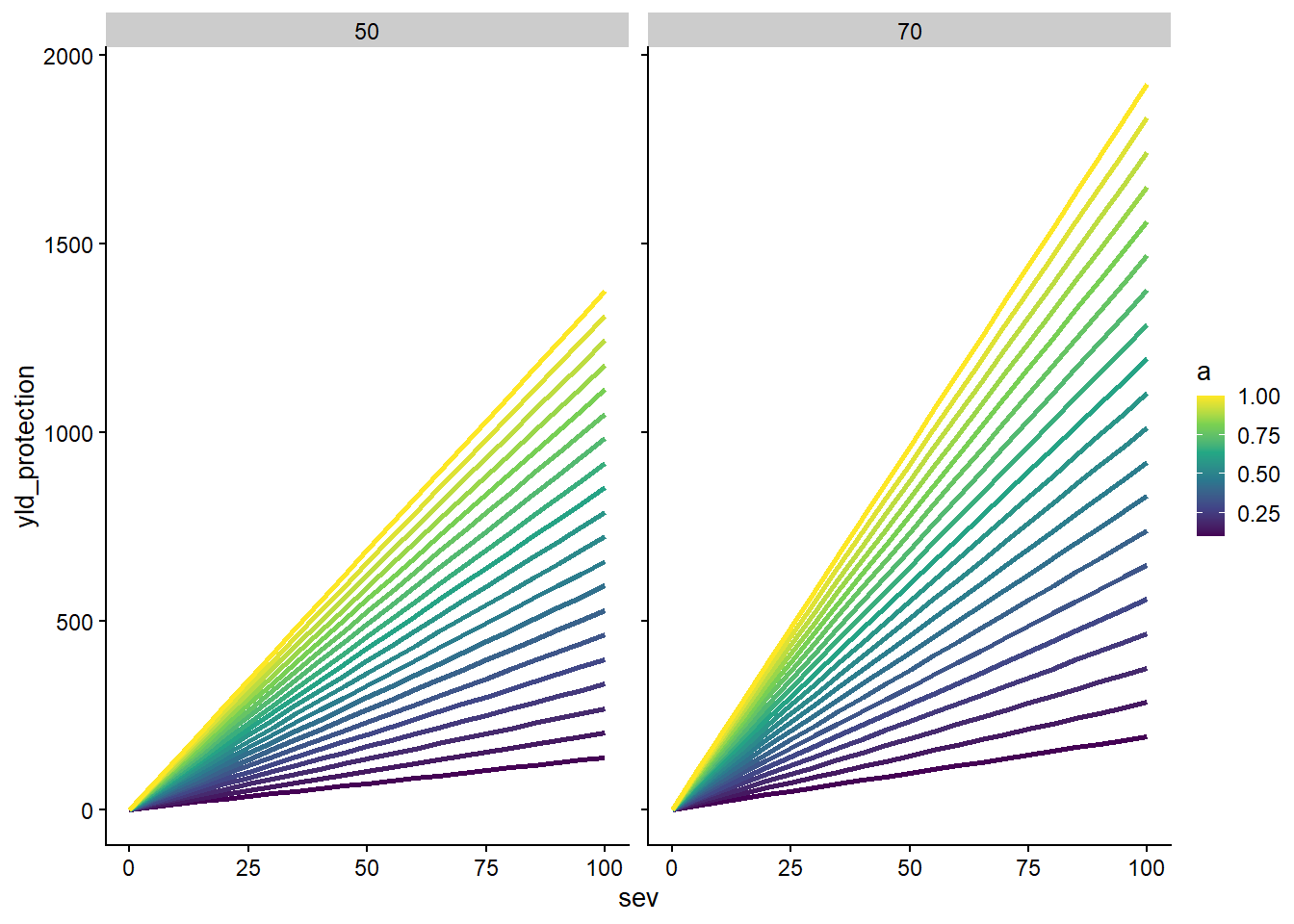
Session Info
sessionInfo()R version 4.4.1 (2024-06-14 ucrt)
Platform: x86_64-w64-mingw32/x64
Running under: Windows 11 x64 (build 26100)
Matrix products: default
locale:
[1] LC_COLLATE=Portuguese_Brazil.utf8 LC_CTYPE=Portuguese_Brazil.utf8
[3] LC_MONETARY=Portuguese_Brazil.utf8 LC_NUMERIC=C
[5] LC_TIME=Portuguese_Brazil.utf8
time zone: America/Sao_Paulo
tzcode source: internal
attached base packages:
[1] stats graphics grDevices datasets utils methods base
other attached packages:
[1] scales_1.3.0 minpack.lm_1.2-4 metafor_4.6-0
[4] numDeriv_2016.8-1.1 metadat_1.2-0 ggthemes_5.1.0
[7] multcomp_1.4-26 TH.data_1.1-2 MASS_7.3-60.2
[10] survival_3.6-4 mvtnorm_1.3-2 emmeans_1.10.6
[13] lmerTest_3.1-3 ggrepel_0.9.6 ggforce_0.4.2
[16] lme4_1.1-35.5 Matrix_1.7-0 lemon_0.4.9
[19] patchwork_1.3.0 cowplot_1.1.3 gsheet_0.4.6
[22] lubridate_1.9.3 forcats_1.0.0 stringr_1.5.1
[25] dplyr_1.1.4 purrr_1.0.2 readr_2.1.5
[28] tidyr_1.3.1 tibble_3.2.1 ggplot2_3.5.1
[31] tidyverse_2.0.0
loaded via a namespace (and not attached):
[1] gridExtra_2.3 gld_2.6.6 sandwich_3.1-1
[4] readxl_1.4.3 rlang_1.1.4 magrittr_2.0.3
[7] e1071_1.7-16 compiler_4.4.1 mgcv_1.9-1
[10] systemfonts_1.1.0 vctrs_0.6.5 quadprog_1.5-8
[13] pkgconfig_2.0.3 fastmap_1.2.0 backports_1.5.0
[16] labeling_0.4.3 utf8_1.2.4 rmarkdown_2.28
[19] tzdb_0.4.0 haven_2.5.4 nloptr_2.1.1
[22] ragg_1.3.3 xfun_0.48 jsonlite_1.8.9
[25] tweenr_2.0.3 broom_1.0.7 parallel_4.4.1
[28] DescTools_0.99.58 R6_2.5.1 stringi_1.8.4
[31] boot_1.3-30 cellranger_1.1.0 estimability_1.5.1
[34] Rcpp_1.0.13 knitr_1.48 zoo_1.8-12
[37] splines_4.4.1 timechange_0.3.0 tidyselect_1.2.1
[40] rstudioapi_0.17.0 yaml_2.3.10 codetools_0.2-20
[43] lattice_0.22-6 plyr_1.8.9 withr_3.0.2
[46] evaluate_1.0.1 proxy_0.4-27 polyclip_1.10-7
[49] ggdist_3.3.2 pillar_1.9.0 BiocManager_1.30.25
[52] renv_1.1.2 distributional_0.5.0 generics_0.1.3
[55] mathjaxr_1.6-0 hms_1.1.3 munsell_0.5.1
[58] rootSolve_1.8.2.4 minqa_1.2.8 class_7.3-22
[61] glue_1.8.0 lmom_3.2 tools_4.4.1
[64] data.table_1.16.2 Exact_3.3 grid_4.4.1
[67] colorspace_2.1-1 nlme_3.1-164 cli_3.6.3
[70] textshaping_0.4.0 fansi_1.0.6 expm_1.0-0
[73] viridisLite_0.4.2 gtable_0.3.5 digest_0.6.37
[76] pbkrtest_0.5.4 farver_2.1.2 htmltools_0.5.8.1
[79] lifecycle_1.0.4 multcompView_0.1-10 httr_1.4.7